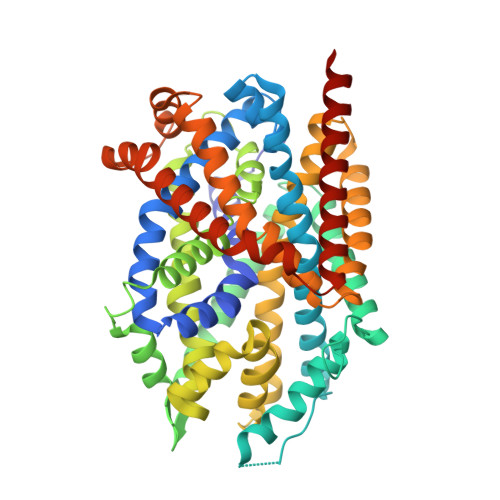
Substrate binds in the S1 site of the F253A mutant of LeuT, a neurotransmitter sodium symporter homologue.
Wang, H., Gouaux, E.(2012) EMBO Rep 13: 861-866
- PubMed: 22836580 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2012.110
- Primary Citation Related Structures:
4FXZ, 4FY0 - PubMed Abstract:
LeuT serves as the model protein for understanding the relationships between structure, mechanism and pharmacology in neurotransmitter sodium symporters (NSSs). At the present time, however, there is a vigorous debate over whether there is a single high-affinity substrate site (S1) located at the original, crystallographically determined substrate site or whether there are two high-affinity substrates sites, one at the primary or S1 site and the other at a second site (S2) located at the base of the extracellular vestibule. In an effort to address the controversy over the number of high-affinity substrate sites in LeuT, one group studied the F253A mutant of LeuT and asserted that in this mutant substrate binds exclusively to the S2 site and that 1 mM clomipramine entirely ablates substrate binding to the S2 site. Here we study the binding of substrate to the F253A mutant of LeuT using ligand binding and X-ray crystallographic methods. Both experimental methods unambiguously show that substrate binds to the S1 site of the F253A mutant and that binding is retained in the presence of 1 mM clomipramine. These studies, in combination with previous work, are consistent with a mechanism for LeuT that involves a single high-affinity substrate binding site.
- Vollum Institute, Oregon Health & Science University, 3181 SW Sam Jackson Park Road, Portland, Oregon 97239, USA.
Organizational Affiliation: