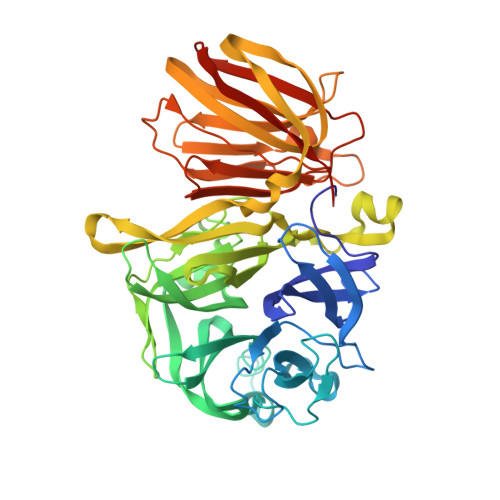
Structural and functional basis for substrate specificity and catalysis of levan fructotransferase.
Park, J., Kim, M.I., Park, Y.D., Shin, I., Cha, J., Kim, C.H., Rhee, S.(2012) J Biological Chem 287: 31233-31241
- PubMed: 22810228
- DOI: https://doi.org/10.1074/jbc.M112.389270
- Primary Citation Related Structures:
4FFF, 4FFG, 4FFH, 4FFI - PubMed Abstract:
Levan is β-2,6-linked polymeric fructose and serves as reserve carbohydrate in some plants and microorganisms. Mobilization of fructose is usually mediated by enzymes such as glycoside hydrolase (GH), typically releasing a monosaccharide as a product. The enzyme levan fructotransferase (LFTase) of the GH32 family catalyzes an intramolecular fructosyl transfer reaction and results in production of cyclic difructose dianhydride, thus exhibiting a novel substrate specificity. The mechanism by which LFTase carries out these functions via the structural fold conserved in the GH32 family is unknown. Here, we report the crystal structure of LFTase from Arthrobacter ureafaciens in apo form, as well as in complexes with sucrose and levanbiose, a difructosacchride with a β-2,6-glycosidic linkage. Despite the similarity of its two-domain structure to members of the GH32 family, LFTase contains an active site that accommodates a difructosaccharide using the -1 and -2 subsites. This feature is unique among GH32 proteins and is facilitated by small side chain residues in the loop region of a catalytic β-propeller N-domain, which is conserved in the LFTase family. An additional oligosaccharide-binding site was also characterized in the β-sandwich C-domain, supporting its role in carbohydrate recognition. Together with functional analysis, our data provide a molecular basis for the catalytic mechanism of LFTase and suggest functional variations from other GH32 family proteins, notwithstanding the conserved structural elements.
- Department of Agricultural Biotechnology, Seoul National University, Seoul 151-921, Korea.
Organizational Affiliation: