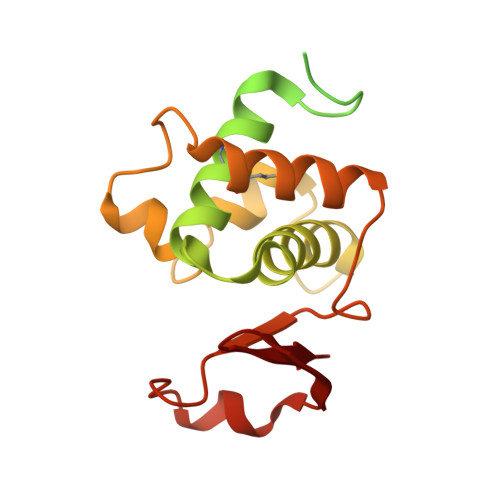
The catalytic domain of the germination-specific lytic transglycosylase SleB from Bacillus anthracis displays a unique active site topology.
Jing, X., Robinson, H.R., Heffron, J.D., Popham, D.L., Schubot, F.D.(2012) Proteins 80: 2469-2475
- PubMed: 22777830 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24140
- Primary Citation Related Structures:
4FET - PubMed Abstract:
Bacillus anthracis produces metabolically inactive spores. Germination of these spores requires germination-specific lytic enzymes (GSLEs) that degrade the unique cortex peptidoglycan to permit resumption of metabolic activity and outgrowth. We report the first crystal structure of the catalytic domain of a GSLE, SleB. The structure revealed a transglycosylase fold with unique active site topology and permitted identification of the catalytic glutamate residue. Moreover, the structure provided insights into the molecular basis for the specificity of the enzyme for muramic-δ-lactam-containing cortex peptidoglycan. The protein also contains a metal-binding site that is positioned directly at the entrance of the substrate-binding cleft.
- Virginia Tech, Department of Biological Sciences, Life Sciences, Blacksburg, Virginia 24061, USA.
Organizational Affiliation: