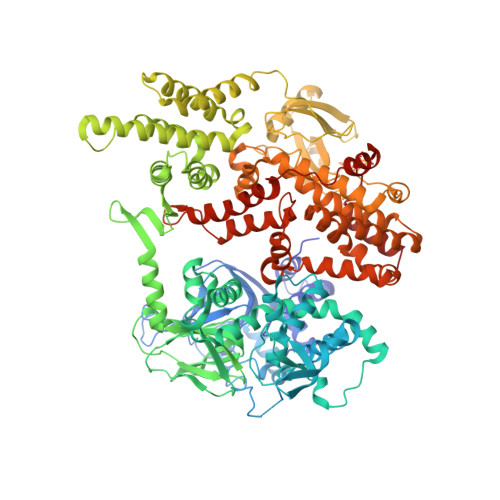
Structural analyses of a constitutively active mutant of exchange protein directly activated by cAMP.
White, M.A., Li, S., Tsalkova, T., Mei, F.C., Liu, T., Woods, V.L., Cheng, X.(2012) PLoS One 7: e49932-e49932
- PubMed: 23189173 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0049932
- Primary Citation Related Structures:
4F7Z - PubMed Abstract:
Exchange proteins directly activated by cAMP (EPACs) are important allosteric regulators of cAMP-mediated signal transduction pathways. To understand the molecular mechanism of EPAC activation, we have combined site-directed mutagenesis, X-ray crystallography, and peptide amide hydrogen/deuterium exchange mass spectrometry (DXMS) to probe the structural and conformational dynamics of EPAC2-F435G, a constitutively active EPAC2 mutant. Our study demonstrates that conformational dynamics plays a critical role in cAMP-induced EPAC activation. A glycine mutation at 435 position shifts the equilibrium of conformational dynamics towards the extended active conformation.
- Sealy Center for Structural Biology and Molecular Biophysics, The University of Texas Medical Branch, Galveston, Texas, United States of America.
Organizational Affiliation: