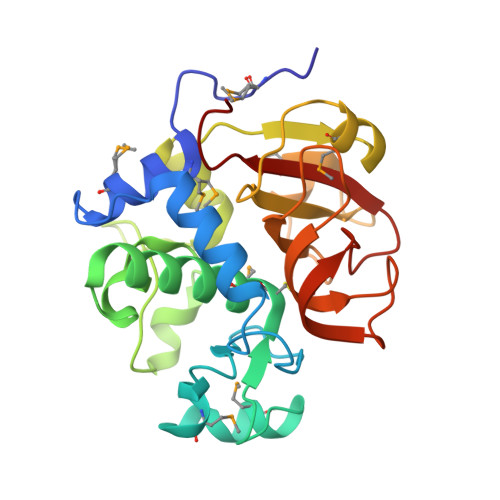
Crystal Structure of an Uncommon Cellulosome-Related Protein Module from Ruminococcus flavefaciens That Resembles Papain-Like Cysteine Peptidases.
Levy-Assaraf, M., Voronov-Goldman, M., Rozman Grinberg, I., Weiserman, G., Shimon, L.J., Jindou, S., Borovok, I., White, B.A., Bayer, E.A., Lamed, R., Frolow, F.(2013) PLoS One 8: e56138-e56138
- PubMed: 23457513 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0056138
- Primary Citation Related Structures:
4EYZ - PubMed Abstract:
Ruminococcus flavefaciens is one of the predominant fiber-degrading bacteria found in the rumen of herbivores. Bioinformatic analysis of the recently sequenced genome indicated that this bacterium produces one of the most intricate cellulosome systems known to date. A distinct ORF, encoding for a multi-modular protein, RflaF_05439, was discovered during mining of the genome sequence. It is composed of two tandem modules of currently undefined function that share 45% identity and a C-terminal X-dockerin modular dyad. Gaining insight into the diversity, architecture and organization of different types of proteins in the cellulosome system is essential for broadening our understanding of a multi-enzyme complex, considered to be one of the most efficient systems for plant cell wall polysaccharide degradation in nature. Following bioinformatic analysis, the second tandem module of RflaF_05439 was cloned and its selenium-labeled derivative was expressed and crystallized. The crystals belong to space group P21 with unit-cell parameters of a = 65.81, b = 60.61, c = 66.13 Å, β = 107.66° and contain two protein molecules in the asymmetric unit. The crystal structure was determined at 1.38-Å resolution by X-ray diffraction using the single-wavelength anomalous dispersion (SAD) method and was refined to Rfactor and Rfree of 0.127 and 0.152 respectively. The protein molecule mainly comprises a β-sheet flanked by short α-helixes, and a globular α-helical domain. The structure was found to be structurally similar to members of the NlpC/P60 superfamily of cysteine peptidases. The 3D structure of the second repeat of the RflaF_05439 enabled us to propose a role for the currently undefined function of this protein. Its putative function as a cysteine peptidase is inferred from in silico structural homology studies. It is therefore apparent that cellulosomes integrate proteins with other functions in addition to the classic well-defined carbohydrate active enzymes.
- Department of Molecular Microbiology and Biotechnology, Tel Aviv University, Tel Aviv, Israel.
Organizational Affiliation: