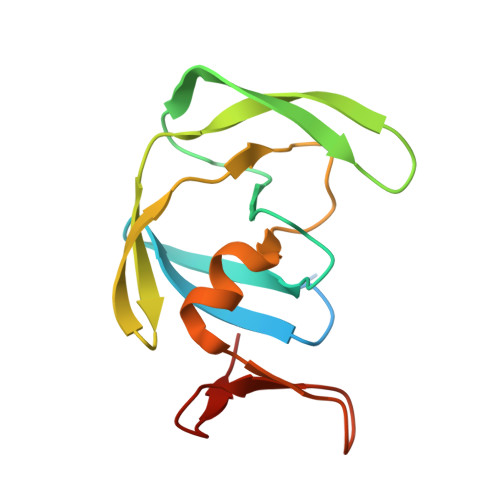
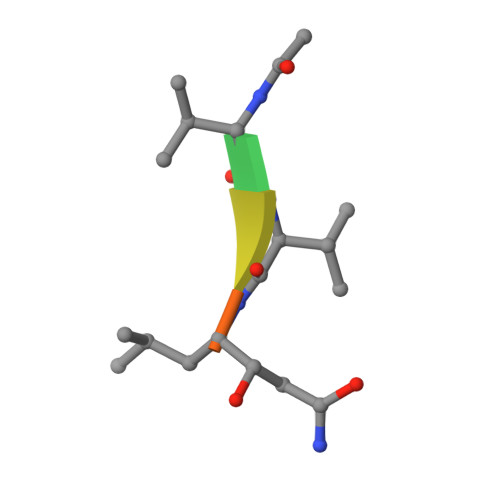
Inhibition of XMRV and HIV-1 proteases by pepstatin A and acetyl-pepstatin.
Matuz, K., Motyan, J., Li, M., Wlodawer, A.(2012) FEBS J 279: 3276-3286
- PubMed: 22804908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1742-4658.2012.08714.x
- Primary Citation Related Structures:
4EXH - PubMed Abstract:
The kinetic properties of two classical inhibitors of aspartic proteases (PRs), pepstatin A and acetyl-pepstatin, were compared in their interactions with HIV-1 and xenotropic murine leukemia virus related virus (XMRV) PRs. Both compounds are substantially weaker inhibitors of XMRV PR than of HIV-1 PR. Previous kinetic and structural studies characterized HIV-1 PR-acetyl-pepstatin and XMRV PR-pepstatin A complexes and suggested dramatically different binding modes. Interaction energies were calculated for the possible binding modes and suggested a strong preference for the one-inhibitor binding mode for HIV-1 PR-acetyl-pepstatin and the two-inhibitor binding mode for XMRV PR-pepstatin A interactions. Comparison of the molecular models suggested that in the case of XMRV PR the relatively unfavorable interactions at S3' and the favorable interactions at S4 and S4' sites with the statine residues may shift the ground state binding towards the two-inhibitor binding mode, whereas the single molecule ground state binding of statines to the HIV-1 PR appear to be more favorable. The preferred single molecular binding to HIV-1 PR allows the formation of the transition state complex, represented by substantially better binding constants. Intriguingly, the crystal structure of the complex of acetyl-pepstatin with XMRV PR has shown a mixed type of binding: the unusual binding mode of two molecules of the inhibitor to the enzyme, in a mode very similar to the previously determined complex with pepstatin A, together with the classical binding mode found for HIV-1 PR. The structure is thus in good agreement with the very similar interaction energies calculated for the two types of binding.
- Department of Biochemistry and Molecular Biology, Faculty of Medicine, University of Debrecen, Hungary.
Organizational Affiliation: