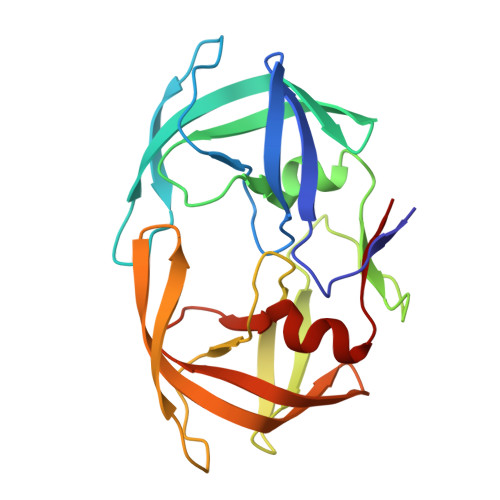
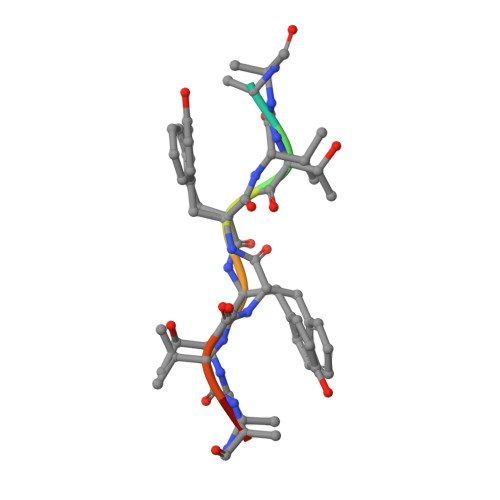
Structural, kinetic, and thermodynamic studies of specificity designed HIV-1 protease.
Alvizo, O., Mittal, S., Mayo, S.L., Schiffer, C.A.(2012) Protein Sci 21: 1029-1041
- PubMed: 22549928
- DOI: https://doi.org/10.1002/pro.2086
- Primary Citation Related Structures:
4EP2, 4EP3, 4EPJ, 4EQ0, 4EQJ - PubMed Abstract:
HIV-1 protease recognizes and cleaves more than 12 different substrates leading to viral maturation. While these substrates share no conserved motif, they are specifically selected for and cleaved by protease during viral life cycle. Drug resistant mutations evolve within the protease that compromise inhibitor binding but allow the continued recognition of all these substrates. While the substrate envelope defines a general shape for substrate recognition, successfully predicting the determinants of substrate binding specificity would provide additional insights into the mechanism of altered molecular recognition in resistant proteases. We designed a variant of HIV protease with altered specificity using positive computational design methods and validated the design using X-ray crystallography and enzyme biochemistry. The engineered variant, Pr3 (A28S/D30F/G48R), was designed to preferentially bind to one out of three of HIV protease's natural substrates; RT-RH over p2-NC and CA-p2. In kinetic assays, RT-RH binding specificity for Pr3 increased threefold compared to the wild-type (WT), which was further confirmed by isothermal titration calorimetry. Crystal structures of WT protease and the designed variant in complex with RT-RH, CA-p2, and p2-NC were determined. Structural analysis of the designed complexes revealed that one of the engineered substitutions (G48R) potentially stabilized heterogeneous flap conformations, thereby facilitating alternate modes of substrate binding. Our results demonstrate that while substrate specificity could be engineered in HIV protease, the structural pliability of protease restricted the propagation of interactions as predicted. These results offer new insights into the plasticity and structural determinants of substrate binding specificity of the HIV-1 protease.
- Division of Biology, Biochemistry and Molecular Biophysics Option, California Institute of Technology, Pasadena, California 91125, USA.
Organizational Affiliation: