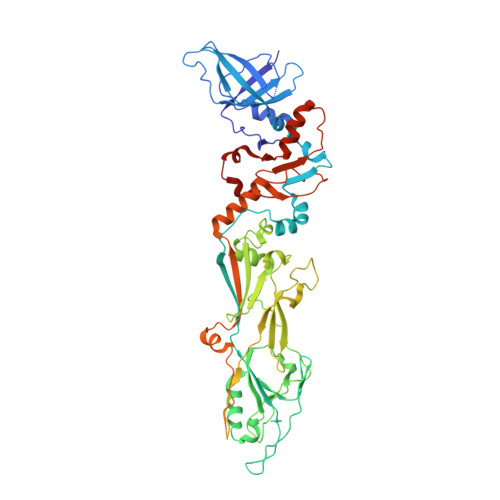
Structural investigations of a Podoviridae streptococcus phage C1, implications for the mechanism of viral entry.
Aksyuk, A.A., Bowman, V.D., Kaufmann, B., Fields, C., Klose, T., Holdaway, H.A., Fischetti, V.A., Rossmann, M.G.(2012) Proc Natl Acad Sci U S A 109: 14001-14006
- PubMed: 22891295
- DOI: https://doi.org/10.1073/pnas.1207730109
- Primary Citation Related Structures:
4EO2, 4EP0 - PubMed Abstract:
The Podoviridae phage C1 was one of the earliest isolated bacteriophages and the first virus documented to be active against streptococci. The icosahedral and asymmetric reconstructions of the virus were calculated using cryo-electron microscopy. The capsid protein has an HK97 fold arranged into a T = 4 icosahedral lattice. The C1 tail is terminated with a ϕ29-like knob, surrounded by a skirt of twelve long appendages with novel morphology. Several C1 structural proteins have been identified, including a candidate for an appendage. The crystal structure of the knob has an N-terminal domain with a fold observed previously in tube forming proteins of Siphoviridae and Myoviridae phages. The structure of C1 suggests the mechanisms by which the virus digests the cell wall and ejects its genome. Although there is little sequence similarity to other phages, conservation of the structural proteins demonstrates a common origin of the head and tail, but more recent evolution of the appendages.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907-2032, USA.
Organizational Affiliation: