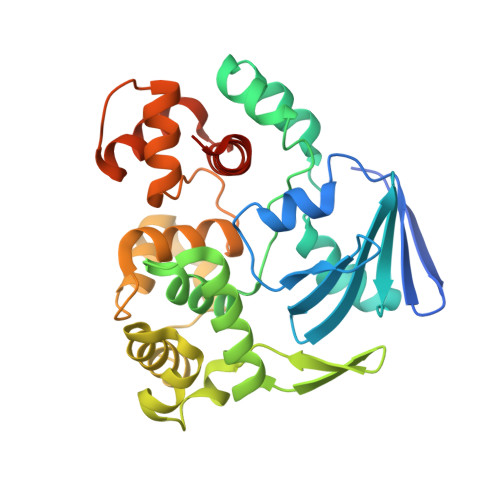
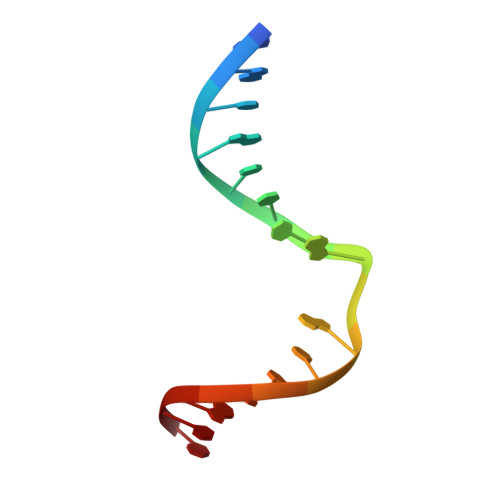
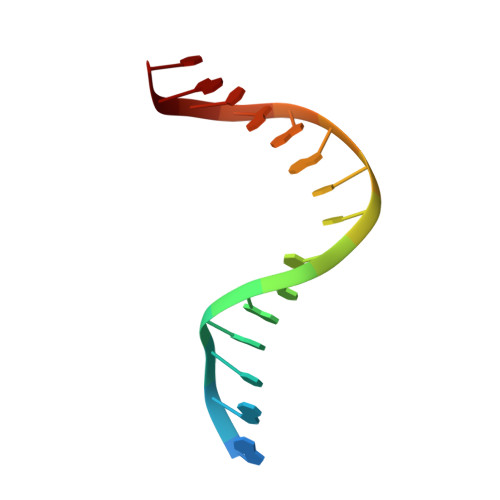
Crystal structures of MBOgg1 in complex with two abasic DNA ligands
Yu, H.J., Yang, M.Z., Zhang, X.E., Bi, L.J., Jiang, T.(2013) J Struct Biol 181: 252-263
- PubMed: 23246782 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2012.12.003
- Primary Citation Related Structures:
4EJY, 4EJZ - PubMed Abstract:
7,8-Dihydro-8-oxoguanine (8-oxoG) is one of the most common oxidative DNA lesions. 8-oxoguanine DNA glycosylases (Oggs) detect and excise 8-oxoG through a multiple-step process. To better understand the basis for estranged base recognition, we have solved the crystal structures of MBOgg1, the 8-oxoguanine DNA glycosylase of Thermoanaerobacter tengcongensis, in complex with DNA containing a tetrahydrofuranyl site (THF, a stable abasic site analog) paired with an estranged cytosine (MBOgg1/DNA(THF:C)) or thymine (MBOgg1/DNA(THF:T)). Different states of THF (extrahelical or intrahelical) are observed in the two complexes of the ASU of MBOgg1/DNA(THF:C) structure. Analyses of their different interaction modes reveal that variable contacts on the 5' region flanking the THF abasic site are correlated with the states of the THF. Comparison of MBOgg1/DNA(THF:T) with MBOgg1/DNA(THF:C) indicates that the non-preferred estranged T may affect MBOgg1's contacts with the 5' flank of the lesion strand. Furthermore, we identified a region in MBOgg1 that is rich in positive charges and interacts with the 5' region flanking the lesion. This region is conserved only in non-eukaryotic Oggs, and additional mutagenesis and biochemical assays reveal that it may contribute to the distinct estranged base specificities between eukaryotic and non-eukaryotic Oggs.
- National Key Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, China.
Organizational Affiliation: