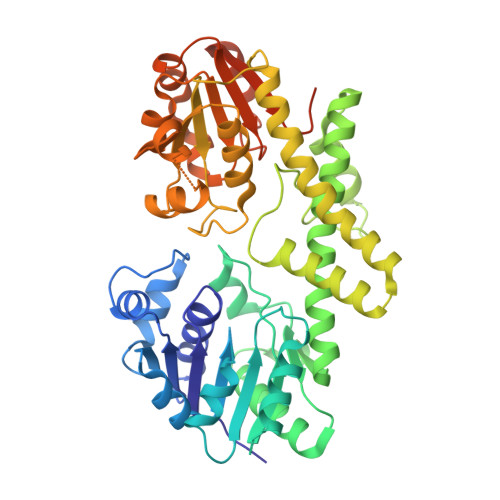
Cofactor binding triggers a molecular switch to allosterically activate human UDP-{alpha}-D-glucose 6-dehydrogenase.
Sennett, N.C., Kadirvelraj, R., Wood, Z.A.(2012) Biochemistry 51: 9364-9374
- PubMed: 23106432
- DOI: https://doi.org/10.1021/bi301067w
- Primary Citation of Related Structures:
4EDF - PubMed Abstract:
Human UDP-α-D-glucose dehydrogenase (hUGDH) catalyzes the NAD(+)-dependent oxidation of UDP-α-D-glucose (UDG) to produce UDP-α-D-glucuronic acid. The oligomeric structure of hUGDH is dynamic and can form two distinct hexameric complexes in solution. The active form of hUGDH consists of dimers that undergo a concentration-dependent association to form a hexamer with 32 symmetry. In the presence of the allosteric feedback inhibitor UDP-α-D-xylose (UDX), hUGDH changes shape to form an inactive, horseshoe-shaped complex. Previous studies have identified the UDX-induced allosteric mechanism that changes the hexameric structure to inhibit the enzyme. Here, we investigate the role of the 32 symmetry hexamer in the catalytic cycle. We engineered a stable hUGDH dimer by introducing a charge-switch substitution (K94E) in the hexamer-building interface (hUGDH(K94E)). The k(cat) of hUGDH(K94E) is ~160-fold lower than that of the wild-type enzyme, suggesting that the hexamer is the catalytically relevant state. We also show that cofactor binding triggers the formation of the 32 symmetry hexamer, but UDG is needed for the stability of the complex. The hUGDH(K94E) crystal structure at 2.08 Å resolution identifies loop(88-110) as the cofactor-responsive allosteric switch that drives hexamer formation; loop(88-110) directly links cofactor binding to the stability of the hexamer-building interface. In the interface, loop(88-110) packs against the Thr131-loop/α6 helix, the allosteric switch that responds to the feedback inhibitor UDX. We also identify a structural element (the S-loop) that explains the indirect stabilization of the hexamer by substrate and supports a sequential, ordered binding of the substrate and cofactor. These observations support a model in which (i) UDG binds to the dimer and stabilizes the S-loop to promote cofactor binding and (ii) cofactor binding orders loop(88-110) to induce formation of the catalytically active hexamer.
- Department of Biochemistry and Molecular Biology, University of Georgia, Athens, Georgia 30602, United States.
Organizational Affiliation: