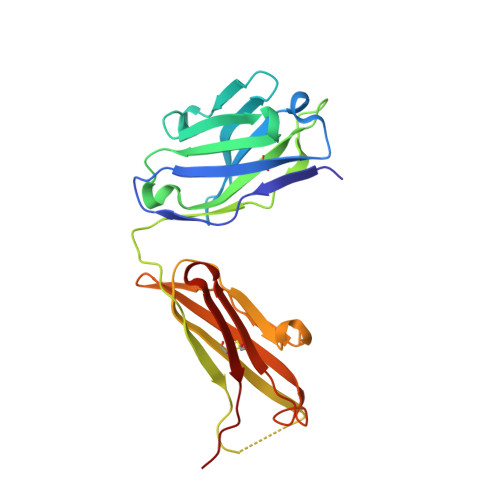
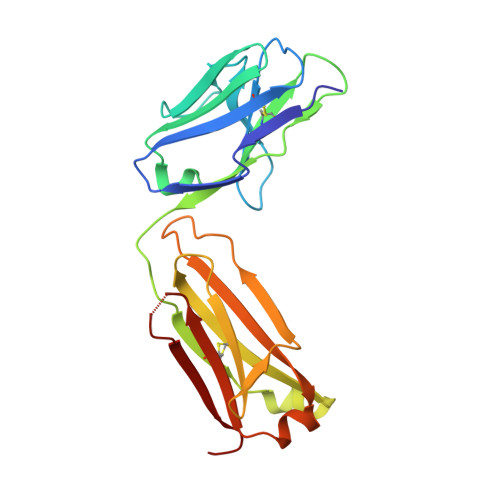
Structural and Biochemical Characterization of the Vaccinia Virus Envelope Protein D8 and Its Recognition by the Antibody LA5.
Matho, M.H., Maybeno, M., Benhnia, M.R., Becker, D., Meng, X., Xiang, Y., Crotty, S., Peters, B., Zajonc, D.M.(2012) J Virol 86: 8050-8058
- PubMed: 22623786 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00836-12
- Primary Citation Related Structures:
4E9O, 4EBQ, 4ETQ - PubMed Abstract:
Smallpox vaccine is considered a gold standard of vaccines, as it is the only one that has led to the complete eradication of an infectious disease from the human population. B cell responses are critical for the protective immunity induced by the vaccine, yet their targeted epitopes recognized in humans remain poorly described. Here we describe the biochemical and structural characterization of one of the immunodominant vaccinia virus (VACV) antigens, D8, and its binding to the monoclonal antibody LA5, which is capable of neutralizing VACV in the presence of complement. The full-length D8 ectodomain was found to form a tetramer. We determined the crystal structure of the LA5 Fab-monomeric D8 complex at a resolution of 2.1 Å, as well as the unliganded structures of D8 and LA5-Fab at resolutions of 1.42 Å and 1.6 Å, respectively. D8 features a carbonic anhydrase (CAH) fold that has evolved to bind to the glycosaminoglycan (GAG) chondroitin sulfate (CS) on host cells. The central positively charged crevice of D8 was predicted to be the CS binding site by automated docking experiments. Furthermore, sequence alignment of various poxvirus D8 orthologs revealed that this crevice is structurally conserved. The D8 epitope is formed by 23 discontinuous residues that are spread across 80% of the D8 protein sequence. Interestingly, LA5 binds with a high-affinity lock-and-key mechanism above this crevice with an unusually large antibody-antigen interface, burying 2,434 Å(2) of protein surface.
- Division of Cell Biology, Division of Vaccine Discovery, La Jolla Institute for Allergy and Immunology, La Jolla, California, USA.
Organizational Affiliation: