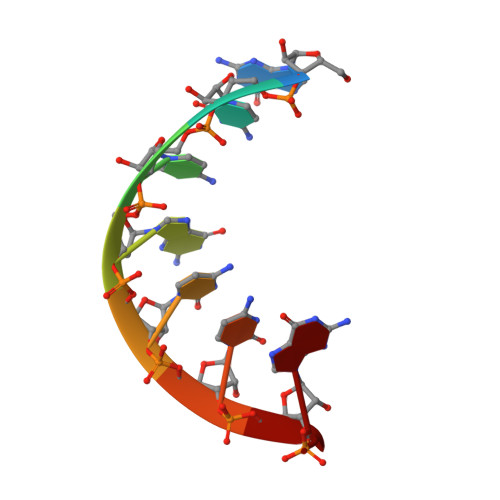
Crystallographic characterization of CCG repeats.
Kiliszek, A., Kierzek, R., Krzyzosiak, W.J., Rypniewski, W.(2012) Nucleic Acids Res 40: 8155-8162
- PubMed: 22718980 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gks557
- Primary Citation Related Structures:
4E58, 4E59 - PubMed Abstract:
CCG repeats are highly over-represented in exons of the human genome. Usually they are located in the 5' UTR but are also abundant in translated sequences. The CCG repeats are associated with three tri-nucleotide repeat disorders: Huntington's disease, myotonic dystrophy type 1 and chromosome X-linked mental retardation (FRAXE). In this study, we present two crystal structures containing double-stranded CCG repeats: one of an RNA in the native form, and one containing LNA nucleotides. Both duplexes form A-helices but with strands slipped in the 5' (native structure) or the 3' direction (LNA-containing structure). As a result, one of two expected C-C pairs is eliminated from the duplex. Each of the three observed C-C pairs interacts differently, forming either one weak H-bond or none. LNA nucleotides have no apparent effect on the helical parameters but the base stacking is increased compared to the native duplex and the distribution of electrostatic potential in the major groove is changed. The CCG crystal structures explain the thermodynamic fragility of CCG runs and throw light on the observation that the MBNL1 protein recognises CCG runs, as well as CUG and CAG, but not the relatively stable CGG repeats.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, Noskowskiego 12/14, 61-704 Poznan, Poland.
Organizational Affiliation: