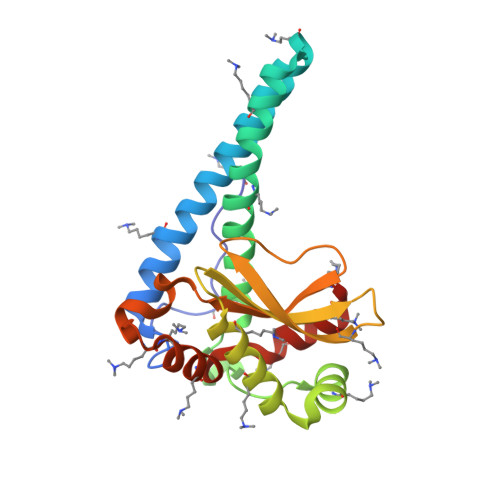
Six-coordinate manganese(3+) in catalysis by yeast manganese superoxide dismutase.
Sheng, Y., Butler Gralla, E., Schumacher, M., Cascio, D., Cabelli, D.E., Selverstone Valentine, J.(2012) Proc Natl Acad Sci U S A 109: 14314-14319
- PubMed: 22908245
- DOI: https://doi.org/10.1073/pnas.1212367109
- Primary Citation Related Structures:
4E4E - PubMed Abstract:
Reduction of superoxide (O2-) by manganese-containing superoxide dismutase occurs through either a "prompt protonation" pathway, or an "inner-sphere" pathway, with the latter leading to formation of an observable Mn-peroxo complex. We recently reported that wild-type (WT) manganese superoxide dismutases (MnSODs) from Saccharomyces cerevisiae and Candida albicans are more gated toward the "prompt protonation" pathway than human and bacterial MnSODs and suggested that this could result from small structural changes in the second coordination sphere of manganese. We report here that substitution of a second-sphere residue, Tyr34, by phenylalanine (Y34F) causes the MnSOD from S. cerevisiae to react exclusively through the "inner-sphere" pathway. At neutral pH, we have a surprising observation that protonation of the Mn-peroxo complex in the mutant yeast enzyme occurs through a fast pathway, leading to a putative six-coordinate Mn(3+) species, which actively oxidizes O2- in the catalytic cycle. Upon increasing pH, the fast pathway is gradually replaced by a slow proton-transfer pathway, leading to the well-characterized five-coordinate Mn(3+). We here propose and compare two hypothetical mechanisms for the mutant yeast enzyme, differing in the structure of the Mn-peroxo complex yet both involving formation of the active six-coordinate Mn(3+) and proton transfer from a second-sphere water molecule, which has substituted for the -OH of Tyr34, to the Mn-peroxo complex. Because WT and the mutant yeast MnSOD both rest in the 2+ state and become six-coordinate when oxidized up from Mn(2+), six-coordinate Mn(3+) species could also actively function in the mechanism of WT yeast MnSODs.
- Department of Chemistry and Biochemistry, Energy-Institute for Genomics and Proteomics, University of California, Los Angeles, CA 90095, USA.
Organizational Affiliation: