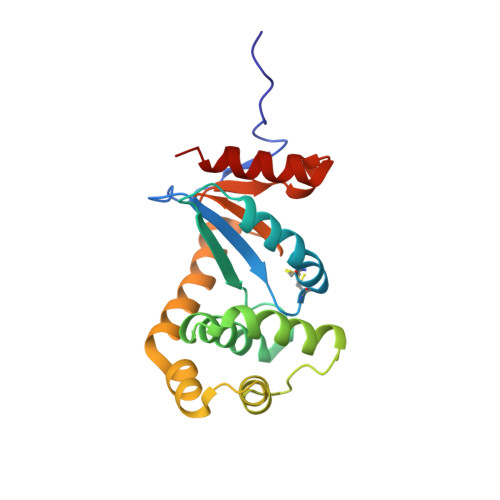
The 1.2 A resolution crystal structure of TcpG, the Vibrio cholerae DsbA disulfide-forming protein required for pilus and cholera-toxin production
Walden, P.M., Heras, B., Chen, K.-E., Halili, M.A., Rimmer, K., Sharma, P., Scanlon, M.J., Martin, J.L.(2012) Acta Crystallogr D Biol Crystallogr 68: 1290-1302
- PubMed: 22993083 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912026388
- Primary Citation Related Structures:
4DVC - PubMed Abstract:
The enzyme TcpG is a periplasmic protein produced by the Gram-negative pathogen Vibrio cholerae. TcpG is essential for the production of ToxR-regulated proteins, including virulence-factor pilus proteins and cholera toxin, and is therefore a target for the development of a new class of anti-virulence drugs. Here, the 1.2 Å resolution crystal structure of TcpG is reported using a cryocooled crystal. This structure is compared with a previous crystal structure determined at 2.1 Å resolution from data measured at room temperature. The new crystal structure is the first DsbA crystal structure to be solved at a sufficiently high resolution to allow the inclusion of refined H atoms in the model. The redox properties of TcpG are also reported, allowing comparison of its oxidoreductase activity with those of other DSB proteins. One of the defining features of the Escherichia coli DsbA enzyme is its destabilizing disulfide, and this is also present in TcpG. The data presented here provide new insights into the structure and redox properties of this enzyme, showing that the binding mode identified between E. coli DsbB and DsbA is likely to be conserved in TcpG and that the β5-α7 loop near the proposed DsbB binding site is flexible, and suggesting that the tense oxidized conformation of TcpG may be the consequence of a short contact at the active site that is induced by disulfide formation and is relieved by reduction.
- Division of Chemistry and Structural Biology, Institute for Molecular Bioscience, University of Queensland, Brisbane, Queensland 4072, Australia.
Organizational Affiliation: