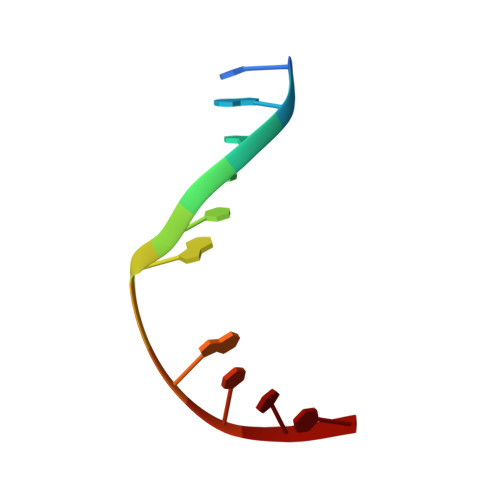
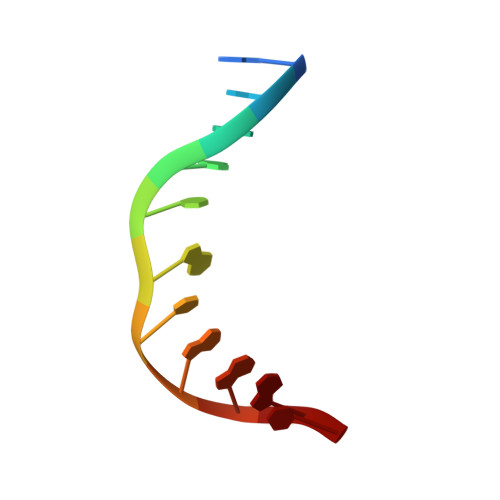
Crystal Structure of Human Methyl-Binding Domain IV Glycosylase Bound to Abasic DNA.
Manvilla, B.A., Maiti, A., Begley, M.C., Toth, E.A., Drohat, A.C.(2012) J Mol Biology 420: 164-175
- PubMed: 22560993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2012.04.028
- Primary Citation Related Structures:
4DK9 - PubMed Abstract:
The mammalian repair protein MBD4 (methyl-CpG-binding domain IV) excises thymine from mutagenic G·T mispairs generated by deamination of 5-methylcytosine (mC), and downstream base excision repair proteins restore a G·C pair. MBD4 is also implicated in active DNA demethylation by initiating base excision repair of G·T mispairs generated by a deaminase enzyme. The question of how mismatch glycosylases attain specificity for excising thymine from G·T, but not A·T, pairs remains largely unresolved. Here, we report a crystal structure of the glycosylase domain of human MBD4 (residues 427-580) bound to DNA containing an abasic nucleotide paired with guanine, providing a glimpse of the enzyme-product complex. The mismatched guanine remains intrahelical, nestled into a recognition pocket. MBD4 provides selective interactions with the mismatched guanine (N1H, N2H(2)) that are not compatible with adenine, which likely confer mismatch specificity. The structure reveals no interactions that would be expected to provide the MBD4 glycosylase domain with specificity for acting at CpG sites. Accordingly, we find modest 1.5- to 2.7-fold reductions in G·T activity upon altering the CpG context. In contrast, 37- to 580-fold effects were observed previously for thymine DNA glycosylase. These findings suggest that specificity of MBD4 for acting at CpG sites depends largely on its methyl-CpG-binding domain, which binds preferably to G·T mispairs in a methylated CpG site. MBD4 glycosylase cannot excise 5-formylcytosine (fC) or 5-carboxylcytosine (caC), intermediates in a Tet (ten eleven translocation)-initiated DNA demethylation pathway. Our structure suggests that MBD4 does not provide the electrostatic interactions needed to excise these oxidized forms of mC.
- Department of Biochemistry and Molecular Biology, University of Maryland School of Medicine, 108 North Greene Street, Baltimore, MD 21201, USA.
Organizational Affiliation: