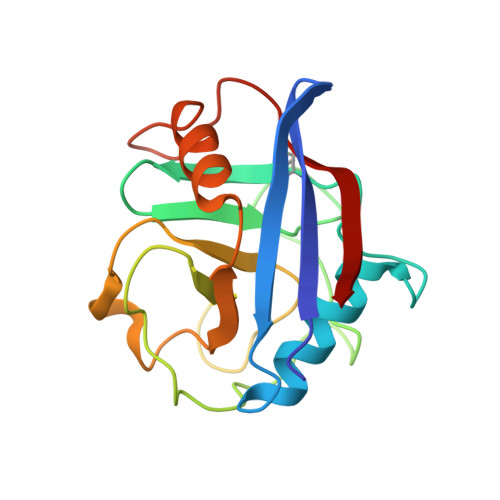
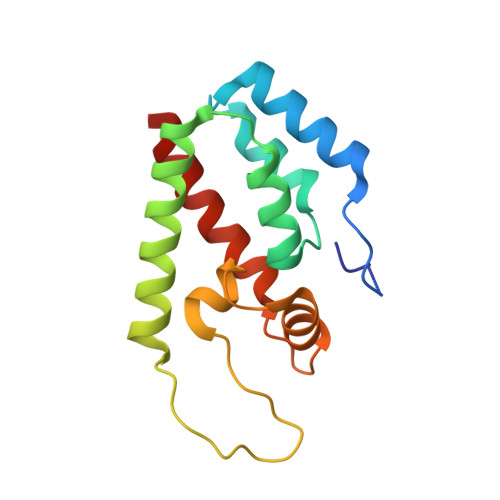
Diverse HIV viruses are targeted by a conformationally dynamic antiviral.
Caines, M.E., Bichel, K., Price, A.J., McEwan, W.A., Towers, G.J., Willett, B.J., Freund, S.M., James, L.C.(2012) Nat Struct Mol Biol 19: 411-416
- PubMed: 22407016 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2253
- Primary Citation Related Structures:
4DGA, 4DGB, 4DGC, 4DGD, 4DGE - PubMed Abstract:
Rhesus macaque TRIMCyp (RhTC) is a potent primate antiviral host protein that inhibits the replication of diverse HIV viruses. Here we show that it has acquired the ability to target multiple viruses by evolving an active site that interconverts between multiple conformations. Mutations that have relieved active site constraints allow RhTC to dynamically sample conformational space, including radically different conformers that target both HIV-1 and HIV-2 viruses. Introduction of a reversible constraint into RhTC allows specificity to be switched between a single conformation specific for HIV-1 and a dynamic ensemble that targets multiple viruses. These results show that conformational diversity can be used to expand the target diversity of innate immune receptors by supplementing their limited genetic variability with variability in protein structure.
- Medical Research Council Laboratory of Molecular Biology, Division of Protein and Nucleic Acid Chemistry, Cambridge, UK.
Organizational Affiliation: