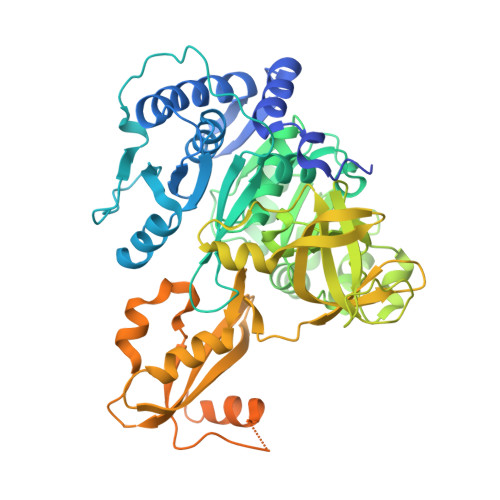
Structure of PA1221, a Nonribosomal Peptide Synthetase Containing Adenylation and Peptidyl Carrier Protein Domains.
Mitchell, C.A., Shi, C., Aldrich, C.C., Gulick, A.M.(2012) Biochemistry 51: 3252-3263
- PubMed: 22452656
- DOI: https://doi.org/10.1021/bi300112e
- Primary Citation Related Structures:
4DG8, 4DG9 - PubMed Abstract:
Many bacteria use large modular enzymes for the synthesis of polyketide and peptide natural products. These multidomain enzymes contain integrated carrier domains that deliver bound substrates to multiple catalytic domains, requiring coordination of these chemical steps. Nonribosomal peptide synthetases (NRPSs) load amino acids onto carrier domains through the activity of an upstream adenylation domain. Our lab recently determined the structure of an engineered two-domain NRPS containing fused adenylation and carrier domains. This structure adopted a domain-swapped dimer that illustrated the interface between these two domains. To continue our investigation, we now examine PA1221, a natural two-domain protein from Pseudomonas aeruginosa. We have determined the amino acid specificity of this new enzyme and used domain specific mutations to demonstrate that loading the downstream carrier domain within a single protein molecule occurs more quickly than loading of a nonfused carrier domain intermolecularly. Finally, we have determined crystal structures of both apo- and holo-PA1221 proteins, the latter using a valine-adenosine vinylsulfonamide inhibitor that traps the adenylation domain-carrier domain interaction. The protein adopts an interface similar to that seen with the prior adenylation domain-carrier protein construct. A comparison of these structures with previous structures of multidomain NRPSs suggests that a large conformational change within the NRPS adenylation domains guides the carrier domain into the active site for thioester formation.
- Hauptman-Woodward Institute and Department of Structural Biology, University at Buffalo, Buffalo, New York 14203, United States.
Organizational Affiliation: