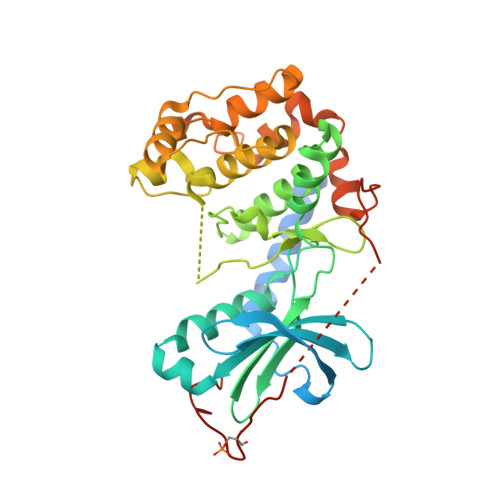
Structural basis for the regulation of protein kinase a by activation loop phosphorylation.
Steichen, J.M., Kuchinskas, M., Keshwani, M.M., Yang, J., Adams, J.A., Taylor, S.S.(2012) J Biological Chem 287: 14672-14680
- PubMed: 22334660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.335091
- Primary Citation Related Structures:
4DFY - PubMed Abstract:
The catalytic subunit of cAMP-dependent protein kinase (PKA) is a member of the AGC group of protein kinases. Whereas PKA has served as a structural model for the protein kinase superfamily, all previous structures of the catalytic subunit contain a phosphorylated activation loop. To understand the structural effects of activation loop phosphorylation at Thr-197 we used a PKA mutant that does not autophosphorylate at Thr-197. The enzyme crystallized in the apo-state, and the structure was solved to 3.0 Å. The N-lobe is rotated by 18° relative to the wild-type apoenzyme, which illustrates that the enzyme likely exists in a wide range of conformations in solution due to the uncoupling of the N- and C-lobes. Several regions of the protein including the activation loop are disordered in the structure, and there are alternate main chain conformations for the magnesium positioning loop and catalytic loop causing a complete loss of hydrogen bonding between these two active site structural elements. These alterations are reflected in a 20-fold decrease in the apparent phosphoryl transfer rate as measured by pre-steady-state kinetic methods.
- Department of Chemistry and Biochemistry, University of California at San Diego, La Jolla, California 92093, USA.
Organizational Affiliation: