Rapid development of piperidine carboxamides as potent and selective anaplastic lymphoma kinase inhibitors.
Bryan, M.C., Whittington, D.A., Doherty, E.M., Falsey, J.R., Cheng, A.C., Emkey, R., Brake, R.L., Lewis, R.T.(2012) J Med Chem 55: 1698-1705
- PubMed: 22263917 Search on PubMed
- DOI: https://doi.org/10.1021/jm201565s
- Primary Citation Related Structures:
4DCE - PubMed Abstract:
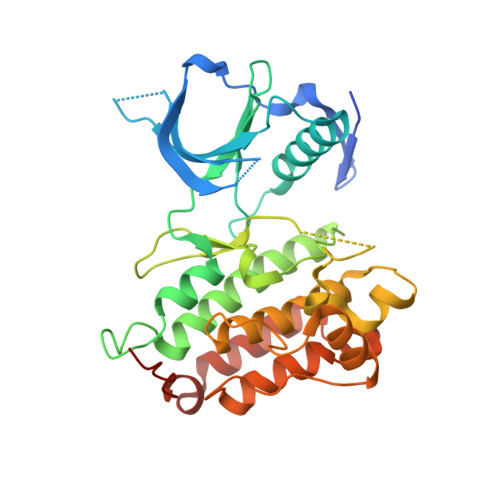
Piperidine carboxamide 1 was identified as a novel inhibitor of anaplastic lymphoma kinase (ALK enzyme assay IC(50) = 0.174 μM) during high throughput screening, with selectivity over the related kinase insulin-like growth factor-1 (IGF1R). The X-ray cocrystal structure of 1 with the ALK kinase domain revealed an unusual DFG-shifted conformation, allowing access to an extended hydrophobic pocket. Structure-activity relationship (SAR) studies were focused on the rapid parallel optimization of both the right- and left-hand side of the molecule, culminating in molecules with improved potency and selectivity over IGF1R.
- Medicinal Chemistry Research Technologies, Amgen Inc., One Amgen Center Drive, Thousand Oaks, California 91320, United States.
Organizational Affiliation: