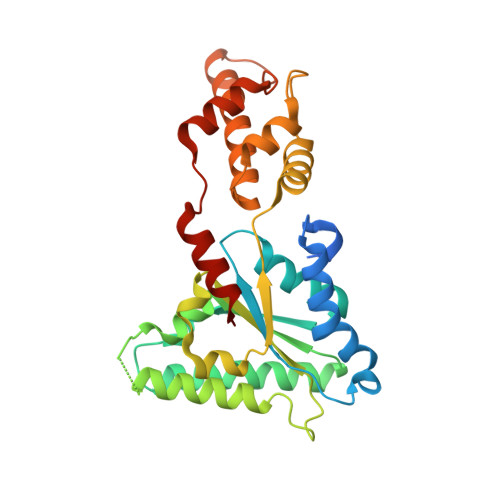
Asymmetric Ring Structure of Vps4 Required for Escrt-III Disassembly.
Caillat, C., Macheboeuf, P., Wu, Y., Mccarthy, A.A., Boeri-Erba, E., Effantin, G., Gottlinger, H.G., Weissenhorn, W., Renesto, P.(2015) Nat Commun 6: 8781
- PubMed: 26632262 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms9781
- Primary Citation Related Structures:
4D80, 4D81, 4D82 - PubMed Abstract:
The vacuolar protein sorting 4 AAA-ATPase (Vps4) recycles endosomal sorting complexes required for transport (ESCRT-III) polymers from cellular membranes. Here we present a 3.6-Å X-ray structure of ring-shaped Vps4 from Metallosphera sedula (MsVps4), seen as an asymmetric pseudohexamer. Conserved key interface residues are shown to be important for MsVps4 assembly, ATPase activity in vitro, ESCRT-III disassembly in vitro and HIV-1 budding. ADP binding leads to conformational changes within the protomer, which might propagate within the ring structure. All ATP-binding sites are accessible and the pseudohexamer binds six ATP with micromolar affinity in vitro. In contrast, ADP occupies one high-affinity and five low-affinity binding sites in vitro, consistent with conformational asymmetry induced on ATP hydrolysis. The structure represents a snapshot of an assembled Vps4 conformation and provides insight into the molecular motions the ring structure undergoes in a concerted action to couple ATP hydrolysis to ESCRT-III substrate disassembly.
- Unit of Virus-Host Cell interactions (UVHCI), University of Grenoble Alpes, F-38042 Grenoble, France.
Organizational Affiliation: