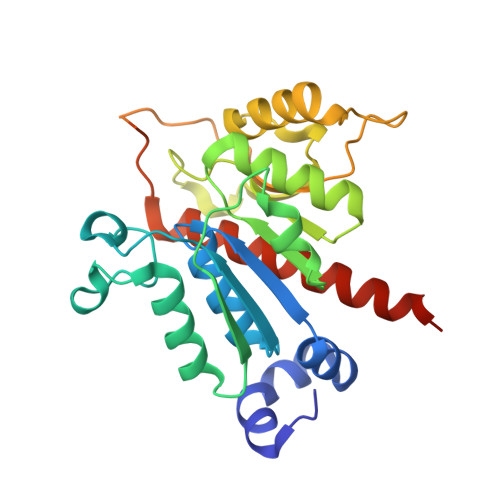
The Crystal Structure and Small-Angle X-Ray Analysis of Csdl/Tcda Reveal a New tRNA Binding Motif in the Moeb/E1 Superfamily.
Lopez-Estepa, M., Arda, A., Savko, M., Round, A., Shepard, W.E., Bruix, M., Coll, M., Fernandez, F.J., Jimenez-Barbero, J., Vega, M.C.(2015) PLoS One 10: 18606
- PubMed: 25897750
- DOI: https://doi.org/10.1371/journal.pone.0118606
- Primary Citation Related Structures:
4D79, 4D7A - PubMed Abstract:
Cyclic N6-threonylcarbamoyladenosine ('cyclic t6A', ct(6)A) is a non-thiolated hypermodification found in transfer RNAs (tRNAs) in bacteria, protists, fungi and plants. In bacteria and yeast cells ct(6)A has been shown to enhance translation fidelity and efficiency of ANN codons by improving the faithful discrimination of aminoacylated tRNAs by the ribosome. To further the understanding of ct(6)A biology we have determined the high-resolution crystal structures of CsdL/TcdA in complex with AMP and ATP, an E1-like activating enzyme from Escherichia coli, which catalyzes the ATP-dependent dehydration of t6A to form ct(6)A. CsdL/TcdA is a dimer whose structural integrity and dimer interface depend critically on strongly bound K+ and Na+ cations. By using biochemical assays and small-angle X-ray scattering we show that CsdL/TcdA can associate with tRNA with a 1:1 stoichiometry and with the proper position and orientation for the cyclization of t6A. Furthermore, we show by nuclear magnetic resonance that CsdL/TcdA engages in transient interactions with CsdA and CsdE, which, in the latter case, involve catalytically important residues. These short-lived interactions may underpin the precise channeling of sulfur atoms from cysteine to CsdL/TcdA as previously characterized. In summary, the combination of structural, biophysical and biochemical methods applied to CsdL/TcdA has afforded a more thorough understanding of how the structure of this E1-like enzyme has been fine tuned to accomplish ct(6)A synthesis on tRNAs while providing support for the notion that CsdA and CsdE are able to functionally interact with CsdL/TcdA.
- Chemical and Physical Biology, Center for Biological Research (CIB-CSIC), Madrid, Spain.
Organizational Affiliation: