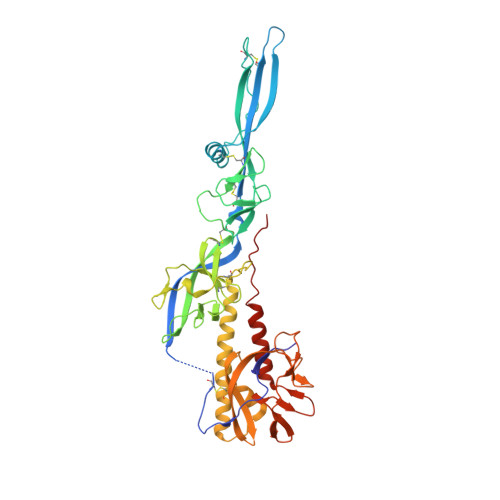
Structure of the Low Ph Conformation of Chandipura Virus G Reveals Important Features in the Evolution of the Vesiculovirus Glycoprotein.
Baquero, E., Albertini, A.A., Raux, H., Buonocore, L., Rose, J.K., Bressanelli, S., Gaudin, Y.(2015) PLoS Pathog 11: 04756
- PubMed: 25803715 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1004756
- Primary Citation Related Structures:
4D6W - PubMed Abstract:
Chandipura virus (CHAV), a member of the vesiculovirus genus, is an emerging human pathogen. As for other rhabdoviruses, CHAV entry into susceptible cells is mediated by its single envelope glycoprotein G which is both involved in receptor recognition and fusion of viral and cellular membranes. Here, we have characterized the fusion properties of CHAV-G. As for vesicular stomatitis virus (VSV, the prototype of the genus) G, fusion is triggered at low pH below 6.5. We have also analyzed the biochemical properties of a soluble form of CHAV-G ectodomain (CHAV-Gth, generated by thermolysin limited-proteolysis of recombinant VSV particles in which the G gene was replaced by that of CHAV). The overall behavior of CHAV-Gth is similar to that previously reported for VSV-Gth. Particularly, CHAV-Gth pre-fusion trimer is not stable in solution and low-pH-induced membrane association of CHAV-Gth is reversible. Furthermore, CHAV-Gth was crystallized in its low pH post-fusion conformation and its structure was determined at 3.6Å resolution. An overall comparison of this structure with the previously reported VSV-Gth post-fusion conformation, shows a high structural similarity as expected from the comparison of primary structure. Among the three domains of G, the pleckstrin homology domain (PHD) appears to be the most divergent and the largest differences are confined to the secondary structure of the major antigenic site of rhabdoviruses. Finally, local differences indicate that CHAV has evolved alternate structural solutions in hinge regions between PH and fusion domains but also distinct pH sensitive switches. Globally the comparison between the post fusion conformation of CHAV and VSV-G highlights several features essential for the protein's function. It also reveals the remarkable plasticity of G in terms of local structures.
- Institute for Integrative Biology of the Cell (I2BC), CEA, CNRS, Université Paris-Sud, Gif-sur-Yvette, France.
Organizational Affiliation: