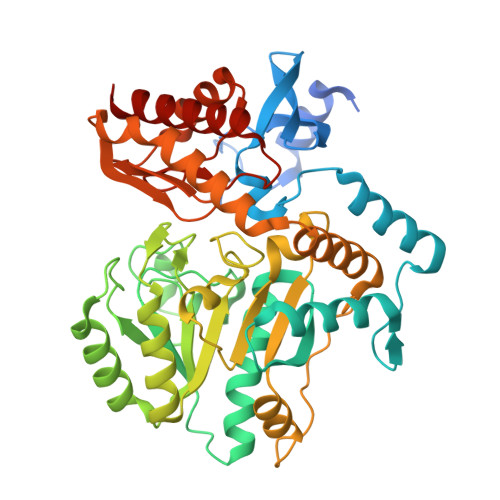
Inhibition of Mycobacterium Tuberculosis Transaminase Bioa by Aryl Hydrazines and Hydrazides.
Dai, R., Wilson, D.J., Geders, T.W., Aldrich, C.C., Finzel, B.C.(2014) Chembiochem 15: 575
- PubMed: 24482078 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cbic.201300748
- Primary Citation Related Structures:
4CXQ, 4CXR, 4MQP, 4MQQ, 4MQR - PubMed Abstract:
7,8-Diaminopelargonic acid synthase (BioA) of Mycobacterium tuberculosis is a recently validated target for therapeutic intervention in the treatment of tuberculosis (TB). Using biophysical fragment screening and structural characterization of compounds, we have identified a potent aryl hydrazine inhibitor of BioA that reversibly modifies the pyridoxal-5'-phosphate (PLP) cofactor, forming a stable quinonoid. Analogous hydrazides also form covalent adducts that can be observed crystallographically but are incapable of inactivating the enzyme. In the X-ray crystal structures, small molecules induce unexpected conformational remodeling in the substrate binding site. We compared these conformational changes to those induced upon binding of the substrate (7-keto-8-aminopelargonic acid), and characterized the inhibition kinetics and the X-ray crystal structures of BioA with the hydrazine compound and analogues to unveil the mechanism of this reversible covalent modification.
- Department of Medicinal Chemistry, University of Minnesota, 308 Harvard St. SE, Minneapolis, MN 55455 (USA).
Organizational Affiliation: