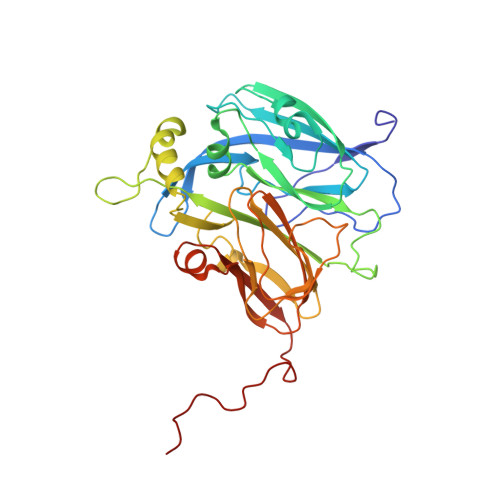
Impact of Residues Remote from the Catalytic Centre on Enzyme Catalysis of Copper Nitrite Reductase.
Leferink, N.G.H., Antonyuk, S.V., Houwman, J.A., Scrutton, N.S., Eady, R.R., Hasnain, S.S.(2014) Nat Commun 5: 4395
- PubMed: 25022223 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms5395
- Primary Citation Related Structures:
4CSP, 4CSZ - PubMed Abstract:
Enzyme mechanisms are often probed by structure-informed point mutations and measurement of their effects on enzymatic properties to test mechanistic hypotheses. In many cases, the challenge is to report on complex, often inter-linked elements of catalysis. Evidence for long-range effects on enzyme mechanism resulting from mutations remains sparse, limiting the design/redesign of synthetic catalysts in a predictable way. Here we show that improving the accessibility of the active site pocket of copper nitrite reductase by mutation of a surface-exposed phenylalanine residue (Phe306), located 12 Å away from the catalytic site type-2 Cu (T2Cu), profoundly affects intra-molecular electron transfer, substrate-binding and catalytic activity. Structures and kinetic studies provide an explanation for the lower affinity for the substrate and the alteration of the rate-limiting step in the reaction. Our results demonstrate that distant residues remote from the active site can have marked effects on enzyme catalysis, by driving mechanistic change through relatively minor structural perturbations.
- 1] Manchester Institute of Biotechnology, Faculty of Life Sciences, University of Manchester, Manchester M1 7DN, UK [2].
Organizational Affiliation: