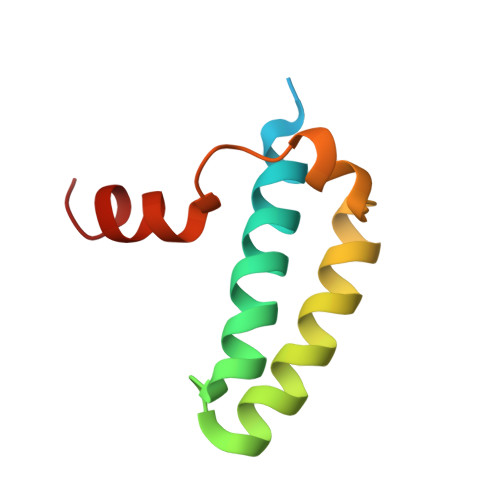
Solution Structure of the Sgta Dimerisation Domain and Investigation of its Interactions with the Ubiquitin-Like Domains of Bag6 and Ubl4A.
Darby, J.F., Krysztofinska, E.M., Simpson, P.J., Simon, A.C., Leznicki, P., Sriskandarajah, N., Bishop, D.S., Hale, L.R., Alfano, C., Conte, M.R., Martinez-Lumbreras, S., Thapaliya, A., High, S., Isaacson, R.L.(2014) PLoS One 9: 11328
- PubMed: 25415308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0113281
- Primary Citation Related Structures:
4CPG - PubMed Abstract:
The BAG6 complex resides in the cytosol and acts as a sorting point to target diverse hydrophobic protein substrates along their appropriate paths, including proteasomal degradation and ER membrane insertion. Composed of a trimeric complex of BAG6, TRC35 and UBL4A, the BAG6 complex is closely associated with SGTA, a co-chaperone from which it can obtain hydrophobic substrates. SGTA consists of an N-terminal dimerisation domain (SGTA_NT), a central tetratricopeptide repeat (TPR) domain, and a glutamine rich region towards the C-terminus. Here we solve a solution structure of the SGTA dimerisation domain and use biophysical techniques to investigate its interaction with two different UBL domains from the BAG6 complex. The SGTA_NT structure is a dimer with a tight hydrophobic interface connecting two sets of four alpha helices. Using a combination of NMR chemical shift perturbation, isothermal titration calorimetry (ITC) and microscale thermophoresis (MST) experiments we have biochemically characterised the interactions of SGTA with components of the BAG6 complex, the ubiquitin-like domain (UBL) containing proteins UBL4A and BAG6. We demonstrate that the UBL domains from UBL4A and BAG6 directly compete for binding to SGTA at the same site. Using a combination of structural and interaction data we have implemented the HADDOCK protein-protein interaction docking tool to generate models of the SGTA-UBL complexes. This atomic level information contributes to our understanding of the way in which hydrophobic proteins have their fate decided by the collaboration between SGTA and the BAG6 complex.
- Department of Chemistry, King's College London, Britannia House, 7 Trinity Street, London, SE1 1DB, United Kingdom.
Organizational Affiliation: