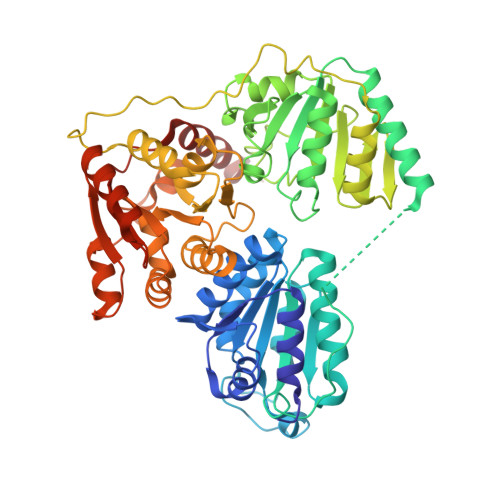
Structure and Functional Characterization of Pyruvate Decarboxylase from Gluconacetobacter Diazotrophicus.
Van Zyl, L.J., Schubert, W., Tuffin, M.I., Cowan, D.A.(2014) BMC Struct Biol 14: 21
- PubMed: 25369873
- DOI: https://doi.org/10.1186/s12900-014-0021-1
- Primary Citation Related Structures:
4COK - PubMed Abstract:
Bacterial pyruvate decarboxylases (PDC) are rare. Their role in ethanol production and in bacterially mediated ethanologenic processes has, however, ensured a continued and growing interest. PDCs from Zymomonas mobilis (ZmPDC), Zymobacter palmae (ZpPDC) and Sarcina ventriculi (SvPDC) have been characterized and ZmPDC has been produced successfully in a range of heterologous hosts. PDCs from the Acetobacteraceae and their role in metabolism have not been characterized to the same extent. Examples include Gluconobacter oxydans (GoPDC), G. diazotrophicus (GdPDC) and Acetobacter pasteutrianus (ApPDC). All of these organisms are of commercial importance. This study reports the kinetic characterization and the crystal structure of a PDC from Gluconacetobacter diazotrophicus (GdPDC). Enzyme kinetic analysis indicates a high affinity for pyruvate (K M 0.06 mM at pH 5), high catalytic efficiencies (1.3 • 10(6) M(-1) • s(-1) at pH 5), pHopt of 5.5 and Topt at 45°C. The enzyme is not thermostable (T½ of 18 minutes at 60°C) and the calculated number of bonds between monomers and dimers do not give clear indications for the relatively lower thermostability compared to other PDCs. The structure is highly similar to those described for Z. mobilis (ZmPDC) and A. pasteurianus PDC (ApPDC) with a rmsd value of 0.57 Å for Cα when comparing GdPDC to that of ApPDC. Indole-3-pyruvate does not serve as a substrate for the enzyme. Structural differences occur in two loci, involving the regions Thr341 to Thr352 and Asn499 to Asp503. This is the first study of the PDC from G. diazotrophicus (PAL5) and lays the groundwork for future research into its role in this endosymbiont. The crystal structure of GdPDC indicates the enzyme to be evolutionarily closely related to homologues from Z. mobilis and A. pasteurianus and suggests strong selective pressure to keep the enzyme characteristics in a narrow range. The pH optimum together with reduced thermostability likely reflect the host organisms niche and conditions under which these properties have been naturally selected for. The lack of activity on indole-3-pyruvate excludes this decarboxylase as the enzyme responsible for indole acetic acid production in G. diazotrophicus.
- Institute for Microbial Biotechnology and Metagenomics (IMBM), University of the Western Cape, Robert Sobukwe Road, Bellville, Cape Town, South Africa. vanzyllj@gmail.com.
Organizational Affiliation: