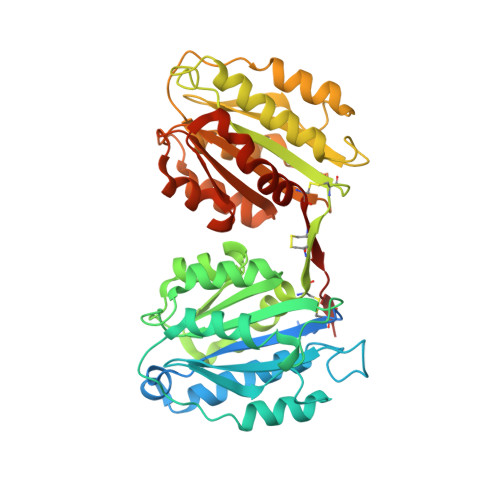
Structural and Functional Features of a Collagen-Binding Matrix Protein from the Mussel Byssus.
Suhre, M.H., Gertz, M., Steegborn, C., Scheibel, T.(2014) Nat Commun 5: 3392
- PubMed: 24569701
- DOI: https://doi.org/10.1038/ncomms4392
- Primary Citation Related Structures:
4CN8, 4CN9, 4CNB - PubMed Abstract:
Blue mussels adhere to surfaces by the byssus, a holdfast structure composed of individual threads representing a collagen fibre reinforced composite. Here, we present the crystal structure and function of one of its matrix proteins, the proximal thread matrix protein 1, which is present in the proximal section of the byssus. The structure reveals two von Willebrand factor type A domains linked by a two-β-stranded linker yielding a novel structural arrangement. In vitro, the protein binds heterologous collagens with high affinity and affects collagen assembly, morphology and arrangement of its fibrils. By providing charged surface clusters as well as insufficiently coordinated metal ions, the proximal thread matrix protein 1 might interconnect other byssal proteins and thereby contribute to the integrity of the byssal threads in vivo. Moreover, the protein could be used for adjusting the mechanical properties of collagen materials, a function likely important in the natural byssus.
- 1] Lehrstuhl Biomaterialien, Fakultät für Ingenieurwissenschaften, Universität Bayreuth, Universitätsstraße 30, Bayreuth 95440, Germany [2].
Organizational Affiliation: