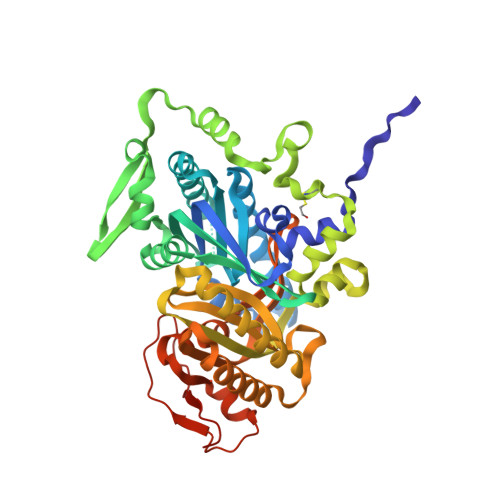
Crystal Structures of Human Soluble Adenylyl Cyclase Reveal Mechanisms of Catalysis and of its Activation Through Bicarbonate.
Kleinboelting, S., Diaz, A., Moniot, S., Van Den Heuvel, J., Weyand, M., Levin, L.R., Buck, J., Steegborn, C.(2014) Proc Natl Acad Sci U S A 111: 3727
- PubMed: 24567411 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1322778111
- Primary Citation Related Structures:
4CLF, 4CLK, 4CLL, 4CLP, 4CLS, 4CLT, 4CLU, 4CLW, 4CLY, 4CLZ, 4CM0, 4CM2 - PubMed Abstract:
cAMP is an evolutionary conserved, prototypic second messenger regulating numerous cellular functions. In mammals, cAMP is synthesized by one of 10 homologous adenylyl cyclases (ACs): nine transmembrane enzymes and one soluble AC (sAC). Among these, only sAC is directly activated by bicarbonate (HCO3(-)); it thereby serves as a cellular sensor for HCO3(-), carbon dioxide (CO2), and pH in physiological functions, such as sperm activation, aqueous humor formation, and metabolic regulation. Here, we describe crystal structures of human sAC catalytic domains in the apo state and in complex with substrate analog, products, and regulators. The activator HCO3(-) binds adjacent to Arg176, which acts as a switch that enables formation of the catalytic cation sites. An anionic inhibitor, 4,4'-diisothiocyanatostilbene-2,2'-disulfonic acid, inhibits sAC through binding to the active site entrance, which blocks HCO3(-) activation through steric hindrance and trapping of the Arg176 side chain. Finally, product complexes reveal small, local rearrangements that facilitate catalysis. Our results provide a molecular mechanism for sAC catalysis and cellular HCO3(-) sensing and a basis for targeting this system with drugs.
- Department of Biochemistry, University of Bayreuth, 95440 Bayreuth, Germany.
Organizational Affiliation: