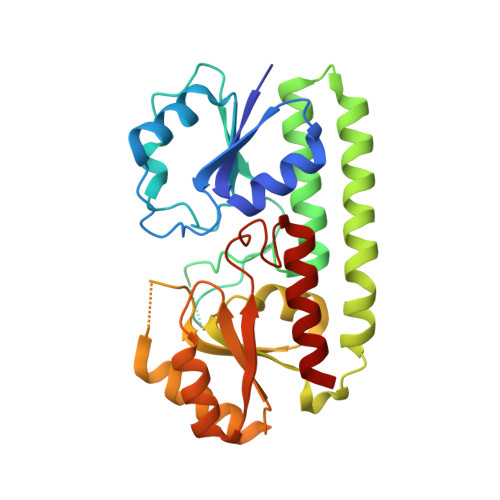
Crystal Structure of a Periplasmic Solute Binding Protein in Metal-Free, Intermediate and Metal-Bound States from Candidatus Liberibacter Asiaticus.
Sharma, N., Selvakumar, P., Bhose, S., Ghosh, D.K., Kumar, P., Sharma, A.K.(2015) J Struct Biol 189: 184
- PubMed: 25641618 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2015.01.012
- Primary Citation Related Structures:
4CL2, 4UDN, 4UDO - PubMed Abstract:
The Znu system, a member of ABC transporter family, is critical for survival and pathogenesis of Candidatus Liberibacter asiaticus (CLA). Two homologues of this system have been identified in CLA. Here, we report high resolution crystal structure of a periplasmic solute binding protein from second of the two gene clusters of Znu system in CLA (CLas-ZnuA2) in metal-free, intermediate and metal-bound states. CLas-ZnuA2 showed maximum sequence identity to the Mn/Fe-specific solute binding proteins (SBPs) of cluster A-I family. The overall fold of CLas-ZnuA2 is similar to the related cluster A-I family SBPs. The sequence and structure analysis revealed the unique features of CLas-ZnuA2. The comparison of CLas-ZnuA2 structure in three states showed that metal binding and release is facilitated by a large displacement along with a change in orientation of the side chain for one of the metal binding residue (His39) flipped away from metal binding site in metal-free form. The crystal structure captured in intermediate state of metal binding revealed the changes in conformation and interaction of the loop hosting His39 during the metal binding. A rigid body movement of C-domain along with partial unfolding of linker helix at its C-terminal during metal binding, as reported for PsaA, was not observed in CLas-ZnuA2. The present results suggest that despite showing maximum sequence identity to the Mn/Fe-specific SBPs, the mechanistic resemblance of CLas-ZnuA2 seems to be closer to Zn-specific SBPs of cluster A-I family.
- Department of Biotechnology, Indian Institute of Technology Roorkee, Roorkee 247 667, India.
Organizational Affiliation: