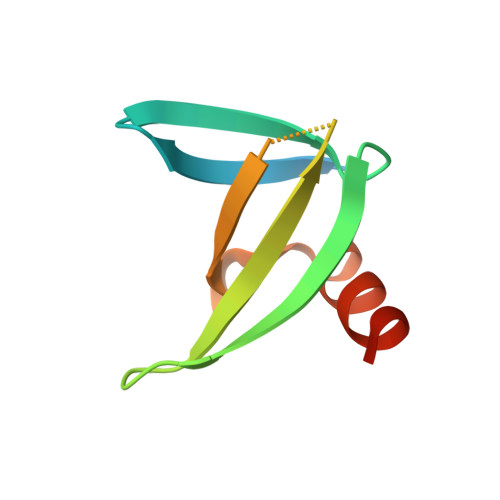
Potent and Specific Inhibition of Glycosidases by Small Artificial Binding Proteins (Affitins)
Correa, A., Pacheco, S., Mechaly, A.E., Obal, G., Behar, G., Mouratou, B., Oppezzo, P., Alzari, P.M., Pecorari, F.(2014) PLoS One 9: 97438
- PubMed: 24823716 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0097438
- Primary Citation Related Structures:
4CJ0, 4CJ1, 4CJ2 - PubMed Abstract:
Glycosidases are associated with various human diseases. The development of efficient and specific inhibitors may provide powerful tools to modulate their activity. However, achieving high selectivity is a major challenge given that glycosidases with different functions can have similar enzymatic mechanisms and active-site architectures. As an alternative approach to small-chemical compounds, proteinaceous inhibitors might provide a better specificity by involving a larger surface area of interaction. We report here the design and characterization of proteinaceous inhibitors that specifically target endoglycosidases representative of the two major mechanistic classes; retaining and inverting glycosidases. These inhibitors consist of artificial affinity proteins, Affitins, selected against the thermophilic CelD from Clostridium thermocellum and lysozyme from hen egg. They were obtained from libraries of Sac7d variants, which involve either the randomization of a surface or the randomization of a surface and an artificially-extended loop. Glycosidase binders exhibited affinities in the nanomolar range with no cross-recognition, with efficient inhibition of lysozyme (Ki = 45 nM) and CelD (Ki = 95 and 111 nM), high expression yields in Escherichia coli, solubility, and thermal stabilities up to 81.1°C. The crystal structures of glycosidase-Affitin complexes validate our library designs. We observed that Affitins prevented substrate access by two modes of binding; covering or penetrating the catalytic site via the extended loop. In addition, Affitins formed salt-bridges with residues essential for enzymatic activity. These results lead us to propose the use of Affitins as versatile selective glycosidase inhibitors and, potentially, as enzymatic inhibitors in general.
- Institut Pasteur de Montevideo, Recombinant Protein Unit, Montevideo, Uruguay; Institut Pasteur, Unité de Microbiologie Structurale, CNRS UMR 3528, Paris, France.
Organizational Affiliation: