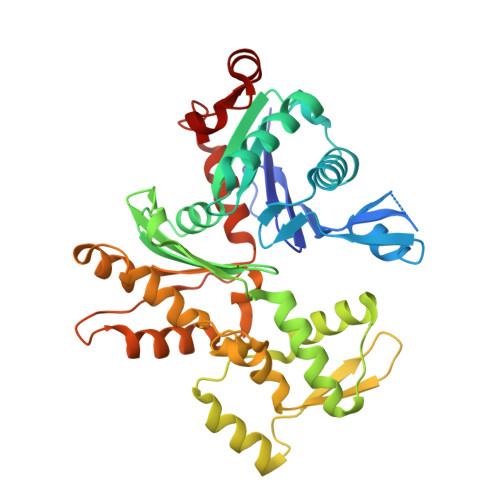
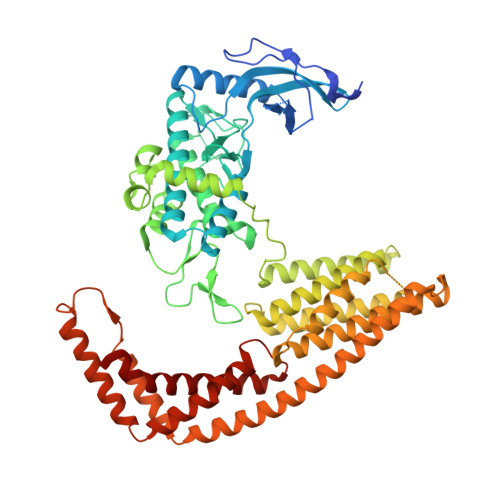
Yersinia Effector Yopo Uses Actin as Bait to Phosphorylate Proteins that Regulate Actin Polymerization.
Lee, W.L., Grimes, J.M., Robinson, R.C.(2015) Nat Struct Mol Biol 22: 248
- PubMed: 25664724
- DOI: https://doi.org/10.1038/nsmb.2964
- Primary Citation Related Structures:
4CI6 - PubMed Abstract:
Pathogenic Yersinia species evade host immune systems through the injection of Yersinia outer proteins (Yops) into phagocytic cells. One Yop, YopO, also known as YpkA, induces actin-filament disruption, impairing phagocytosis. Here we describe the X-ray structure of Yersinia enterocolitica YopO in complex with actin, which reveals that YopO binds to an actin monomer in a manner that blocks polymerization yet allows the bound actin to interact with host actin-regulating proteins. SILAC-MS and biochemical analyses confirm that actin-polymerization regulators such as VASP, EVL, WASP, gelsolin and the formin diaphanous 1 are directly sequestered and phosphorylated by YopO through formation of ternary complexes with actin. This leads to a model in which YopO at the membrane sequesters actin from polymerization while using the bound actin as bait to recruit, phosphorylate and misregulate host actin-regulating proteins to disrupt phagocytosis.
- 1] Institute of Molecular and Cell Biology, Agency for Science, Technology and Research (A*STAR), Singapore. [2] Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, UK.
Organizational Affiliation: