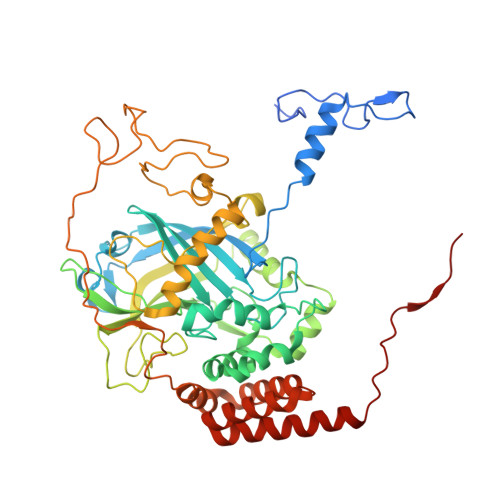
Structure of the Mono-Functional Heme Catalase Dr1998 from Deinococcus Radiodurans
Borges, P.T., Frazao, C., Miranda, C.S., Carrondo, M.A., Romao, C.V.(2014) FEBS J 281: 4138
- PubMed: 24975828 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12895
- Primary Citation Related Structures:
4CAB - PubMed Abstract:
Deinococcus radiodurans is an aerobic organism with the ability to survive under conditions of high radiation doses or desiccation. As part of its protection system against oxidative stress, this bacterium encodes three monofunctional catalases. The DR1998 catalase belongs to clade 1, and is present at high levels under normal growth conditions. The crystals of DR1998 diffracted very weakly, and the merged diffraction data showed an R sym of 0.308. Its crystal structure was determined and refined to 2.6 Å. The four molecules present in the asymmetric unit form, by crystallographic symmetry, two homotetramers with 222 point-group symmetry. The overall structure of DR1998 is similar to that of other monofunctional catalases, showing higher structural homology with the catalase structures of clade 1. Each monomer shows the typical catalase fold, and contains one heme b in the active site. The heme is coordinated by the proximal ligand Tyr369, and on the heme distal side the essential His81 and Asn159 are hydrogen-bonded to a water molecule. A 25-Å-long channel is the main channel connecting the active site to the external surface. This channel starts with a hydrophobic region from the catalytic heme site, which is followed by a hydrophilic region that begins on Asp139 and expands up to the protein surface. Apart from this channel, an alternative channel, also near the heme active site, is presented and discussed. Coordinates and structure factors have been deposited in the Protein Data Bank in Europe under accession code 4CAB.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Oeiras, Portugal.
Organizational Affiliation: