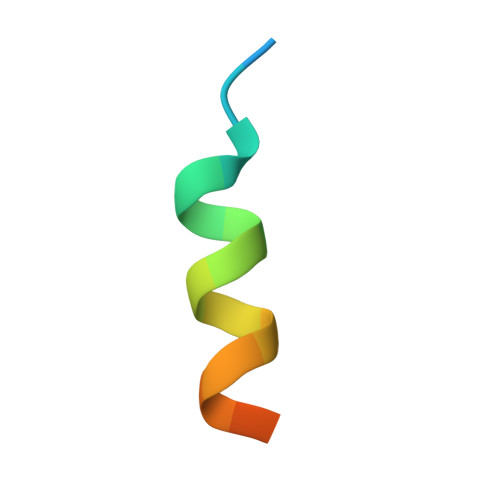
A Ctf4 Trimer Couples the Cmg Helicase to DNA Polymerase a in the Eukaryotic Replisome
Simon, A.C., Zhou, J.C., Perera, R.L., Vandeursen, F., Evrin, C., Ivanova, M.E., Kilkenny, M.L., Renault, L., Kjaer, S., Matak-Vinkovic, D., Labib, K., Costa, A., Pellegrini, L.(2014) Nature 510: 293
- PubMed: 24805245 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature13234
- Primary Citation Related Structures:
4C8H, 4C8S, 4C93, 4C95 - PubMed Abstract:
Efficient duplication of the genome requires the concerted action of helicase and DNA polymerases at replication forks to avoid stalling of the replication machinery and consequent genomic instability. In eukaryotes, the physical coupling between helicase and DNA polymerases remains poorly understood. Here we define the molecular mechanism by which the yeast Ctf4 protein links the Cdc45-MCM-GINS (CMG) DNA helicase to DNA polymerase α (Pol α) within the replisome. We use X-ray crystallography and electron microscopy to show that Ctf4 self-associates in a constitutive disk-shaped trimer. Trimerization depends on a β-propeller domain in the carboxy-terminal half of the protein, which is fused to a helical extension that protrudes from one face of the trimeric disk. Critically, Pol α and the CMG helicase share a common mechanism of interaction with Ctf4. We show that the amino-terminal tails of the catalytic subunit of Pol α and the Sld5 subunit of GINS contain a conserved Ctf4-binding motif that docks onto the exposed helical extension of a Ctf4 protomer within the trimer. Accordingly, we demonstrate that one Ctf4 trimer can support binding of up to three partner proteins, including the simultaneous association with both Pol α and GINS. Our findings indicate that Ctf4 can couple two molecules of Pol α to one CMG helicase within the replisome, providing a new model for lagging-strand synthesis in eukaryotes that resembles the emerging model for the simpler replisome of Escherichia coli. The ability of Ctf4 to act as a platform for multivalent interactions illustrates a mechanism for the concurrent recruitment of factors that act together at the fork.
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, UK.
Organizational Affiliation: