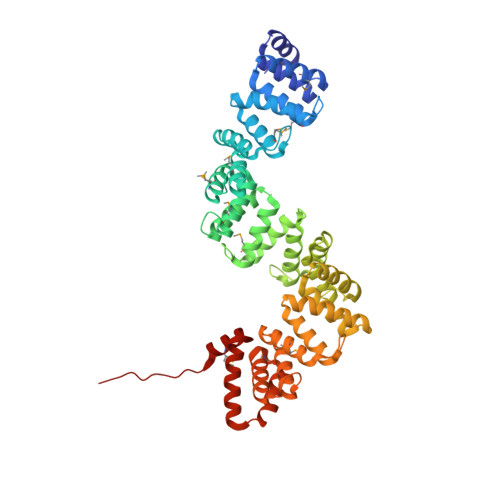
Crystal Structure of C5321: A Protective Antigen Present in Uropathogenic Escherichia Coli Strains Displaying an Slr Fold.
Urosev, D., Ferrer-Navarro, M., Pastorello, I., Cartocci, E., Costenaro, L., Zhulenkovs, D., Marechal, J., Leonchiks, A., Reverter, D., Serino, L., Soriani, M., Daura, X.(2013) BMC Struct Biol 13: 19
- PubMed: 24099525 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-13-19
- Primary Citation Related Structures:
4BWR - PubMed Abstract:
Increasing rates of antimicrobial resistance among uropathogens led, among other efforts, to the application of subtractive reverse vaccinology for the identification of antigens present in extraintestinal pathogenic E. coli (ExPEC) strains but absent or variable in non-pathogenic strains, in a quest for a broadly protective Escherichia coli vaccine. The protein coded by locus c5321 from CFT073 E. coli was identified as one of nine potential vaccine candidates against ExPEC and was able to confer protection with an efficacy of 33% in a mouse model of sepsis. c5321 (known also as EsiB) lacks functional annotation and structurally belongs to the Sel1-like repeat (SLR) family. Herein, as part of the general characterization of this potential antigen, we have focused on its structural properties. We report the 1.74 Å-resolution crystal structure of c5321 from CFT073 E. coli determined by Se-Met SAD phasing. The structure is composed of 11 SLR units in a topological organisation that highly resembles that found in HcpC from Helicobacter pylori, with the main difference residing in how the super-helical fold is stabilised. The stabilising effect of disulfide bridges in HcpC is replaced in c5321 by a strengthening of the inter-repeat hydrophobic core. A metal-ion binding site, uncharacteristic of SLR proteins, is detected between SLR units 3 and 4 in the region of the inter-repeat hydrophobic core. Crystal contacts are observed between the C-terminal tail of one molecule and the C-terminal amphipathic groove of a neighbouring one, resembling interactions between ligand and proteins containing tetratricopeptide-like repeats. The structure of antigen c5321 presents a mode of stabilization of the SLR fold different from that observed in close homologs of known structure. The location of the metal-ion binding site and the observed crystal contacts suggest a potential role in regulation of conformational flexibility and interaction with yet unidentified target proteins, respectively. These findings open new perspectives in both antigen design and for the identification of a functional role for this protective antigen.
- Institute of Biotechnology and Biomedicine, Universitat Autònoma de Barcelona, Bellaterra 08193, Spain. xavier.daura@uab.cat.
Organizational Affiliation: