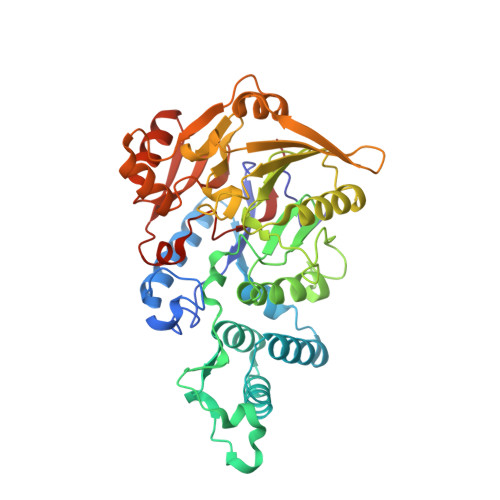
Structure-informed design of an enzymatically inactive vaccine component for group A Streptococcus.
Henningham, A., Ericsson, D.J., Langer, K., Casey, L.W., Jovcevski, B., Chhatwal, G.S., Aquilina, J.A., Batzloff, M.R., Kobe, B., Walker, M.J.(2013) mBio 4
- PubMed: 23919999 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.00509-13
- Primary Citation Related Structures:
4BOF - PubMed Abstract:
Streptococcus pyogenes (group A Streptococcus [GAS]) causes ~700 million human infections/year, resulting in >500,000 deaths. There is no commercial GAS vaccine available. The GAS surface protein arginine deiminase (ADI) protects mice against a lethal challenge. ADI is an enzyme that converts arginine to citrulline and ammonia. Administration of a GAS vaccine preparation containing wild-type ADI, a protein with inherent enzymatic activity, may present a safety risk. In an approach intended to maximize the vaccine safety of GAS ADI, X-ray crystallography and structural immunogenic epitope mapping were used to inform vaccine design. This study aimed to knock out ADI enzyme activity without disrupting the three-dimensional structure or the recognition of immunogenic epitopes. We determined the crystal structure of ADI at 2.5 Å resolution and used it to select a number of amino acid residues for mutagenesis to alanine (D166, E220, H275, D277, and C401). Each mutant protein displayed abrogated activity, and three of the mutant proteins (those with the D166A, H275A, and D277A mutations) possessed a secondary structure and oligomerization state equivalent to those of the wild type, produced high-titer antisera, and avoided disruption of B-cell epitopes of ADI. In addition, antisera raised against the D166A and D277A mutant proteins bound to the GAS cell surface. The inactivated D166A and D277A mutant ADIs are ideal for inclusion in a GAS vaccine preparation. There is no human ortholog of ADI, and we confirm that despite limited structural similarity in the active-site region to human peptidyl ADI 4 (PAD4), ADI does not functionally mimic PAD4 and antiserum raised against GAS ADI does not recognize human PAD4. We present an example of structural biology informing human vaccine design. We previously showed that the administration of the enzyme arginine deiminase (ADI) to mice protected the mice against infection with multiple GAS serotypes. In this study, we determined the structure of GAS ADI and used this information to improve the vaccine safety of GAS ADI. Catalytically inactive mutant forms of ADI retained structure, recognition by antisera, and immunogenic epitopes, rendering them ideal for inclusion in GAS vaccine preparations. This example of structural biology informing vaccine design may underpin the formulation of a safe and efficacious GAS vaccine.
- School of Chemistry and Molecular Biosciences, Australian Infectious Diseases Research Centre, University of Queensland, St. Lucia, Qld., Australia.
Organizational Affiliation: