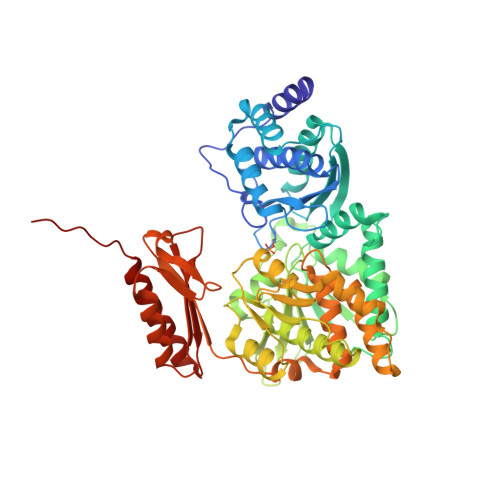
Genetic and Structural Validation of Aspergillus Fumigatus N-Acetylphosphoglucosamine Mutase as an Antifungal Target.
Fang, W., Du, T., Raimi, O.G., Hurtado Guerrero, R., Marino, K., Ibrahim, A.F.M., Albarbarawi, O., Ferguson, M.A.J., Jin, C., Van Aalten, D.M.F.(2013) Biosci Rep 33: 63
- PubMed: 23844980
- DOI: https://doi.org/10.1042/BSR20130053
- Primary Citation Related Structures:
4BJU - PubMed Abstract:
Aspergillus fumigatus is the causative agent of IA (invasive aspergillosis) in immunocompromised patients. It possesses a cell wall composed of chitin, glucan and galactomannan, polymeric carbohydrates synthesized by processive glycosyltransferases from intracellular sugar nucleotide donors. Here we demonstrate that A. fumigatus possesses an active AfAGM1 (A. fumigatus N-acetylphosphoglucosamine mutase), a key enzyme in the biosynthesis of UDP (uridine diphosphate)-GlcNAc (N-acetylglucosamine), the nucleotide sugar donor for chitin synthesis. A conditional agm1 mutant revealed the gene to be essential. Reduced expression of agm1 resulted in retarded cell growth and altered cell wall ultrastructure and composition. The crystal structure of AfAGM1 revealed an amino acid change in the active site compared with the human enzyme, which could be exploitable in the design of selective inhibitors. AfAGM1 inhibitors were discovered by high-throughput screening, inhibiting the enzyme with IC50s in the low μM range. Together, these data provide a platform for the future development of AfAGM1 inhibitors with antifungal activity.
- *Division of Molecular Microbiology, University of Dundee, DD1 5EH, Scotland, U.K.
Organizational Affiliation: