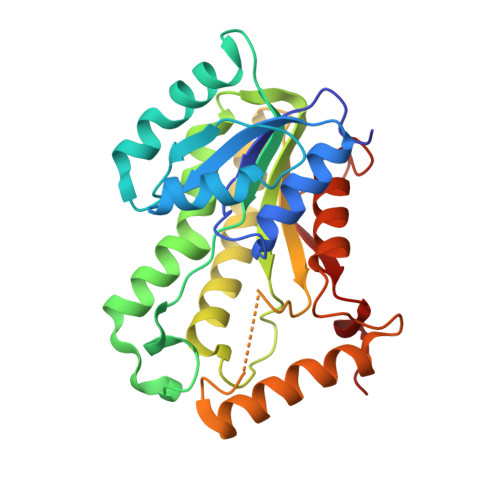
Pyridomycin Bridges the Nadh and Substrate Binding Pockets of the Enoyl Reductase Inha
Hartkoorn, R., Pojer, F., Read, J.A., Gingell, H., Neres, J., Horlacher, O., Altmann, K.H., Cole, S.(2014) Nat Chem Biol 10: 96
- PubMed: 24292073 Search on PubMed
- DOI: https://doi.org/10.1038/nchembio.1405
- Primary Citation Related Structures:
4BGE, 4BGI, 4BII - PubMed Abstract:
Pyridomycin, a natural product with potent antituberculosis activity, inhibits a major drug target, the InhA enoyl reductase. Here, we unveil the co-crystal structure and unique ability of pyridomycin to block both the NADH cofactor- and lipid substrate-binding pockets of InhA. This is to our knowledge a first-of-a-kind binding mode that discloses a new means of InhA inhibition. Proof-of-principle studies show how structure-assisted drug design can improve the activity of new pyridomycin derivatives.
- Ecole Polytechnique Fédérale de Lausanne, Global Health Institute, Lausanne, Switzerland.
Organizational Affiliation: