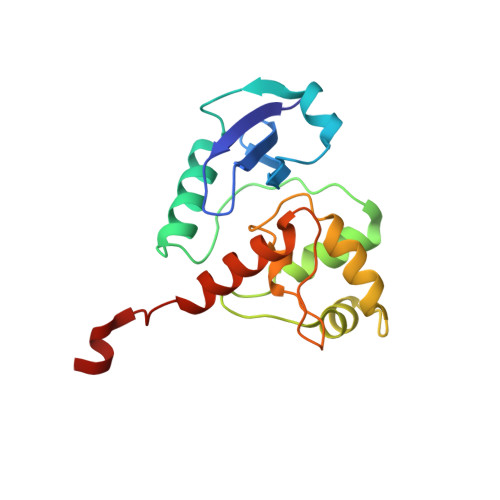
Biochemical and Structural Studies of the Mycobacterium Tuberculosis O6-Methylguanine Methyltransferase and Mutated Variants.
Miggiano, R., Casazza, V., Garavaglia, S., Ciaramella, M., Perugino, G., Rizzi, M., Rossi, F.(2013) J Bacteriol 195: 2728
- PubMed: 23564173 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.02298-12
- Primary Citation Related Structures:
4BHB, 4BHC - PubMed Abstract:
Mycobacterium tuberculosis displays remarkable genetic stability despite continuous exposure to the hostile environment represented by the host's infected macrophages. Similarly to other organisms, M. tuberculosis possesses multiple systems to counteract the harmful potential of DNA alkylation. In particular, the suicidal enzyme O(6)-methylguanine-DNA methyltransferase (OGT) is responsible for the direct repair of O(6)-alkylguanine in double-stranded DNA and is therefore supposed to play a central role in protecting the mycobacterial genome from the risk of G · C-to-A · T transition mutations. Notably, a number of geographically widely distributed M. tuberculosis strains shows nonsynonymous single-nucleotide polymorphisms in their OGT-encoding gene, leading to amino acid substitutions at position 15 (T15S) or position 37 (R37L) of the N-terminal domain of the corresponding protein. However, the role of these mutations in M. tuberculosis pathogenesis is unknown. We describe here the in vitro characterization of M. tuberculosis OGT (MtOGT) and of two point-mutated versions of the protein mimicking the naturally occurring ones, revealing that both mutated proteins are impaired in their activity as a consequence of their lower affinity for alkylated DNA than the wild-type protein. The analysis of the crystal structures of MtOGT and MtOGT-R37L confirms the high level of structural conservation of members of this protein family and provides clues to an understanding of the molecular bases for the reduced affinity for the natural substrate displayed by mutated MtOGT. Our in vitro results could contribute to validate the inferred participation of mutated OGTs in M. tuberculosis phylogeny and biology.
- Dipartimento di Scienze del Farmaco, University of Piemonte Orientale A Avogadro, Novara, Italy.
Organizational Affiliation: