Discovery and Optimization of Pyrimidone Indoline Amide Pi3Kbeta Inhibitors for the Treatment of Phosphatase and Tensin Homologue (Pten)-Deficient Cancers.
Certal, V., Carry, J.B., Halley, F., Virone-Oddos, A., Thompson, F., Filoche-Romme, B., El-Ahmad, Y., Karlsson, A., Charrier, V., Delorme, C., Rak, A., Abecassis, P., Amara, C., Vincent, L., Bonnevaux, H., Nicolas, J., Mathieu, M., Bertrand, T., Marquette, J., Michot, N., Benard, T., Perrin, M., Lemaitre, O., Guerif, S., Perron, S., Monget, S., Gruss-Leleu, F., Doerflinger, G., Guizani, H., Brollo, M., Delbarre, L., Bertin, L., Richepin, P., Loyau, V., Garcia-Echeverria, C., Lengauer, C., Schio, L.(2014) J Med Chem 57: 903
- PubMed: 24387221 Search on PubMed
- DOI: https://doi.org/10.1021/jm401642q
- Primary Citation Related Structures:
4BFR - PubMed Abstract:
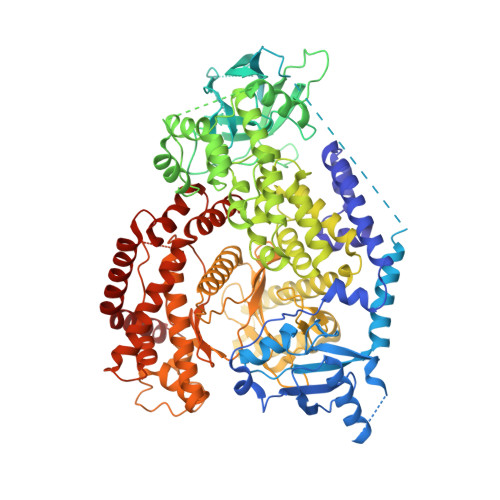
Compelling molecular biology publications have reported the implication of phosphoinositide kinase PI3Kβ in PTEN-deficient cell line growth and proliferation. These findings supported a scientific rationale for the development of PI3Kβ-specific inhibitors for the treatment of PTEN-deficient cancers. This paper describes the discovery of 2-[2-(2,3-dihydro-indol-1-yl)-2-oxo-ethyl]-6-morpholin-4-yl-3H-pyrimidin-4-one (7) and the optimization of this new series of active and selective pyrimidone indoline amide PI3Kβ inhibitors. 2-[2-(2-Methyl-2,3-dihydro-indol-1-yl)-2-oxo-ethyl]-6-morpholin-4-yl-3H-pyrimidin-4-one (28), identified following a carefully designed methyl scan, displayed improved physicochemical and in vitro pharmacokinetic properties. Structural biology efforts enabled the acquisition of the first X-ray cocrystal structure of p110β with the selective inhibitor compound 28 bound to the ATP site. The nonplanar binding mode described herein is consistent with observed structure-activity relationship for the series. Compound 28 demonstrated significant in vivo activity in a UACC-62 xenograft model in mice, warranting further preclinical investigation. Following successful development, compound 28 entered phase I/Ib clinical trial in patients with advanced cancer.
- Oncology Drug Discovery, §Structure Design Informatics, and Structural Biology, #Drug Disposition and Safety (DSAR), †Protein Production,⊥Pharmaceutical Sciences, ∥Analytical Sciences, Sanofi , 13, quai Jules Guesde, 94403 Vitry-sur-Seine, France.
Organizational Affiliation: