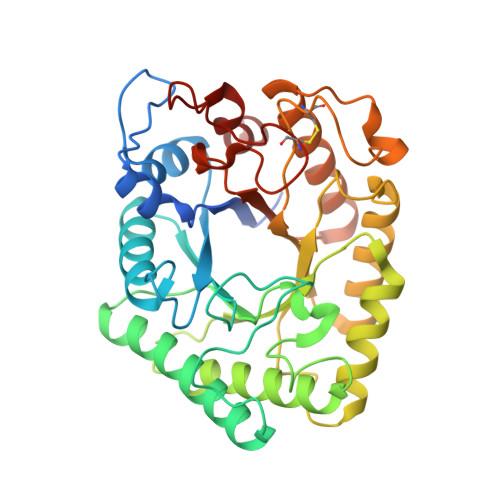
The Binding of Zinc Ions to Emericella Nidulans Endo-[Beta]-1,4-Galactanase is Essential for Crystal Formation
Otten, H., Michalak, M., Mikkelsen, J.D., Larsen, S.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 850
- PubMed: 23908026
- DOI: https://doi.org/10.1107/S1744309113019714
- Primary Citation Related Structures:
4BF7 - PubMed Abstract:
A novel Emericella nidulans endo-β-1,4-galactanase (EnGAL) demonstrates a strong capacity to generate high levels of very potent prebiotic oligosaccharides from potato pulp, a by-product of the agricultural potato-starch industry. EnGAL belongs to glycoside hydrolase family 53 and shows high (72.5%) sequence identity to an endo-β-1,4-galactanase from Aspergillus aculeatus. Diffraction data extending to 2.0 Å resolution were collected from a crystal of EnGAL grown from conditions containing 0.2 M zinc acetate. The crystal structure showed a high similarity between EnGAL and other endo-β-1,4-galactanases belonging to GH53. It also revealed 15 zinc ions bound to the protein, one of which is located in the active site, where it is coordinated by residues Glu136 and Glu246 which comprise the catalytic machinery. The majority of the zinc ions are located on the surface of the enzyme, in some cases with side chains from two different molecules as ligands, thus explaining why the presence of zinc ions was essential for crystallization.
- Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: