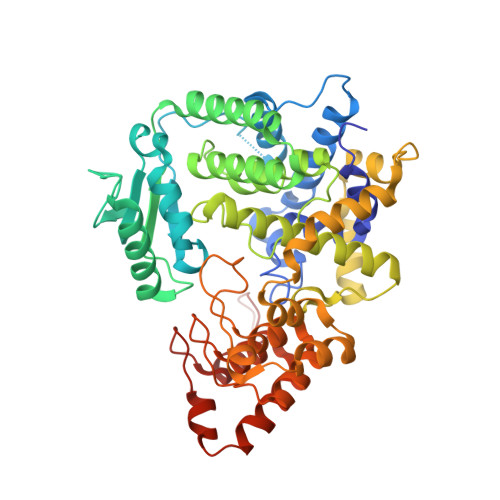
Structure of the Legionella Effector Ankx Reveals the Mechanism of Phosphocholine Transfer by the Fic Domain.
Campanacci, V., Mukherjee, S., Roy, C.R., Cherfils, J.(2013) EMBO J 32: 1469
- PubMed: 23572077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2013.82
- Primary Citation Related Structures:
4BEP, 4BER, 4BES, 4BET - PubMed Abstract:
The FIC motif and the eukaryotic-like ankyrin repeats are found in many bacterial type IV effectors, yet little is known about how these domains enable bacteria to modulate host cell functions. Bacterial FIC domains typically bind ATP and transfer adenosine monophosphate moiety onto target proteins. The ankyrin repeat-containing protein AnkX encoded by the intracellular pathogen Legionella pneumophila is unique in that its FIC domain binds to CDP-choline and transfers a phosphocholine residue onto proteins in the Rab1 GTPase family. By determining the structures of unbound AnkX and AnkX with bound CDP-choline, CMP/phosphocholine and CMP, we demonstrate that the orientation of substrate binding in relation to the catalytic FIC motif enables this protein to function as a phosphocholinating enzyme rather than a nucleotidyl transferase. Additionally, the structure reveals that the ankyrin repeats mediate scaffolding interactions that resemble those found in protein-protein interactions, but are unprecedented in intramolecular interactions. Together with phosphocholination experiments, our structures unify a general phosphoryl transferase mechanism common to all FIC enzymes that should be conserved from bacteria to human.
- Laboratoire d'Enzymologie et Biochimie Structurales, Centre de Recherche de Gif, CNRS, Gif-sur-Yvette, France.
Organizational Affiliation: