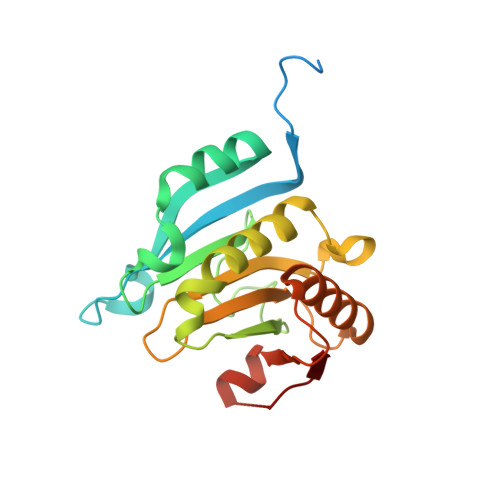
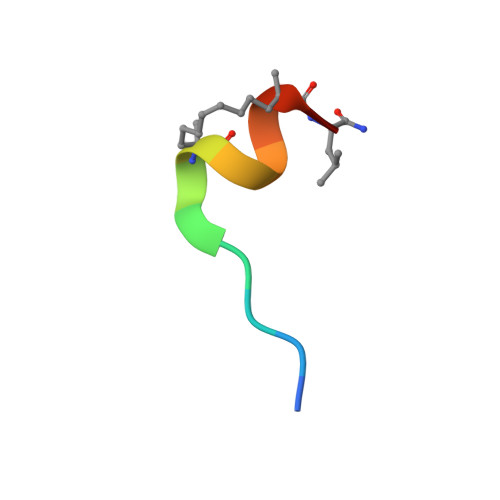
Rational Optimization of Conformational Effects Induced by Hydrocarbon Staples in Peptides and Their Binding Interfaces.
Lama, D., Quah, S.T., Verma, C.S., Lakshminarayanan, R., Beuerman, R.W., Lane, D.P., Brown, C.J.(2013) Sci Rep 3: 3451
- PubMed: 24336354
- DOI: https://doi.org/10.1038/srep03451
- Primary Citation of Related Structures:
4BEA - PubMed Abstract:
eIF4E is frequently over-expressed in different cancers and causes increased translation of oncogenic proteins via deregulated cap-dependent translation. Inhibitors of the eIF4E:eIF4G interactions represent an approach that would normalize cap-dependent translation. Stapled peptides represent an emerging class of therapeutics that can target protein: protein interactions. We present here molecular dynamics simulations for a set of rationally designed stapled peptides in solution and in complex with eIF4E, supported with biophysical and crystallographic data. Clustering of the simulated structures revealed the favoured conformational states of the stapled peptides in their bound or free forms in solution. Identifying these populations has allowed us to design peptides with improved affinities by introducing mutations into the peptide sequence to alter their conformational distributions. These studies emphasise the effects that engineered mutations have on the conformations of free and bound peptides, and illustrate that both states must be considered in efforts to attain high affinity binding.
- 1] Bioinformatics Institute, A*STAR (Agency for Science, Technology and Research), 30 Biopolis Street, #07-01 Matrix, Singapore 138671 [2].
Organizational Affiliation: