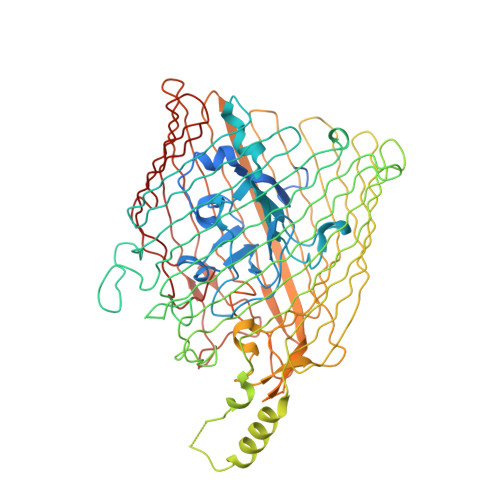
Use of a Molecular Decoy to Segregate Transport from Antigenicity in the Frpb Iron Transporter from Neisseria Meningitidis.
Saleem, M., Prince, S.M., Rigby, S.E.J., Imran, M., Patel, H., Chan, H., Sanders, H., Maiden, M.C.J., Feavers, I.M., Derrick, J.P.(2013) PLoS One 8: 56746
- PubMed: 23457610 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0056746
- Primary Citation Related Structures:
4AIP, 4AIQ, 4B7O - PubMed Abstract:
FrpB is an outer membrane transporter from Neisseria meningitidis, the causative agent of meningococcal meningitis. It is a member of the TonB-dependent transporter (TBDT) family and is responsible for iron uptake into the periplasm. FrpB is subject to a high degree of antigenic variation, principally through a region of hypervariable sequence exposed at the cell surface. From the crystal structures of two FrpB antigenic variants, we identify a bound ferric ion within the structure which induces structural changes on binding which are consistent with it being the transported substrate. Binding experiments, followed by elemental analysis, verified that FrpB binds Fe(3+) with high affinity. EPR spectra of the bound Fe(3+) ion confirmed that its chemical environment was consistent with that observed in the crystal structure. Fe(3+) binding was reduced or abolished on mutation of the Fe(3+)-chelating residues. FrpB orthologs were identified in other Gram-negative bacteria which showed absolute conservation of the coordinating residues, suggesting the existence of a specific TBDT sub-family dedicated to the transport of Fe(3+). The region of antigenic hypervariability lies in a separate, external sub-domain, whose structure is conserved in both the F3-3 and F5-1 variants, despite their sequence divergence. We conclude that the antigenic sub-domain has arisen separately as a result of immune selection pressure to distract the immune response from the primary transport function. This would enable FrpB to function as a transporter independently of antibody binding, by using the antigenic sub-domain as a 'molecular decoy' to distract immune surveillance.
- Michael Smith Building, Faculty of Life Sciences, University of Manchester, Oxford Road, Manchester, United Kingdom.
Organizational Affiliation: