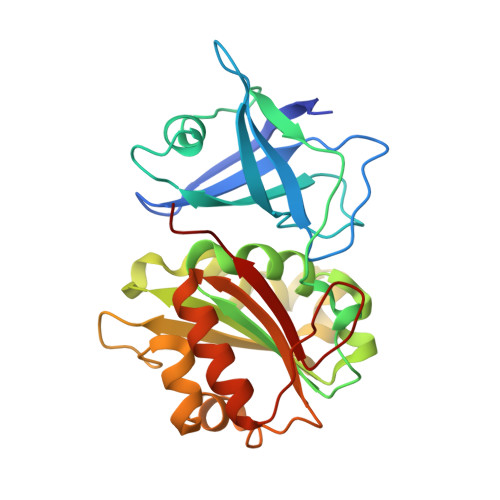
Crystal Structure of Fad-Containing Ferredoxin- Nadp Reductase from Xanthomonas Axonopodis Pv. Citri
Tondo, M.L., Hurtado-Guerrero, R., Sanchez-Azqueta, A., Ceccarelli, E.A., Medina, M., Orellano, E.G., Martinez-Julvez, M.(2013) Biomed Res Int 2013: 06572
- PubMed: 23984418 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1155/2013/906572
- Primary Citation Related Structures:
4B4D - PubMed Abstract:
We have solved the structure of ferredoxin-NADP(H) reductase, FPR, from the plant pathogen Xanthomonas axonopodis pv. citri, responsible for citrus canker, at a resolution of 1.5 Å. This structure reveals differences in the mobility of specific loops when compared to other FPRs, probably unrelated to the hydride transfer process, which contributes to explaining the structural and functional divergence between the subclass I FPRs. Interactions of the C-terminus of the enzyme with the phosphoadenosine of the cofactor FAD limit its mobility, thus affecting the entrance of nicotinamide into the active site. This structure opens the possibility of rationally designing drugs against the X. axonopodis pv. citri phytopathogen.
- Molecular Biology Division, Instituto de Biología Molecular y Celular de Rosario, IBR, CONICET, Facultad de Ciencias Bioquímicas y Farmacéuticas, Universidad Nacional de Rosario, Suipacha 531, S2002LRK Rosario, Argentina.
Organizational Affiliation: