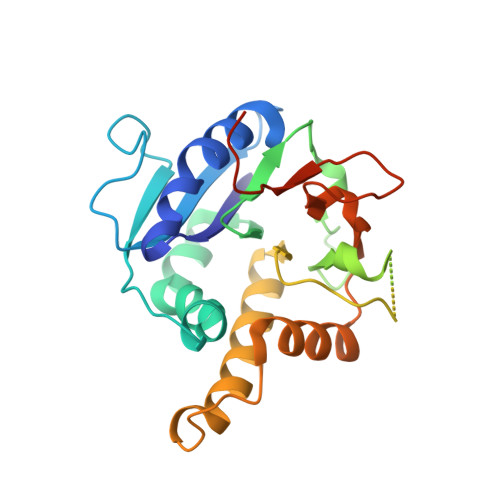
Structure of a Metal-Independent Bacterial Glycosyltransferase that Catalyzes the Synthesis of Histo-Blood Group a Antigen
Thiyagarajan, N., Pham, T.T.K., Stinson, B., Sundriyal, A., Tumbale, P., Lizotte-Waniewski, M., Brew, K., Acharya, K.R.(2012) Sci Rep 2: 940
- PubMed: 23230506
- DOI: https://doi.org/10.1038/srep00940
- Primary Citation Related Structures:
4AYJ, 4AYL - PubMed Abstract:
Histo-blood group antigens (HBGAs) are a source of antigenic variation between individuals that modulates resistance and susceptibility to pathogens and is a barrier to the spread of enveloped viruses. HBGAs are also produced by a few prokaryotes where they are synthesized by glycosyltransferases (GTs) related to human HBGA synthases. Here we report the first structure of a bacterial GT of this family, from an intestinal resident, Bacteroides ovatus. Unlike its mammalian homologues and other GTs with similar folds, this protein lacks a metal-binding Asp-X-Asp motif and is fully active in the absence of divalent metal ions, yet is strikingly similar in structure and in its interactions with substrates to structurally characterized mammalian metal-dependent mammalian homologues. This shows how an apparently major divergence in catalytic properties can be accommodated by minor structural adjustments and illustrates the structural underpinnings of horizontal transfer of a functional gene from prokaryotes to vertebrates.
- Department of Biology and Biochemistry, University of Bath , Claverton Down, Bath BA2 7AY, UK.
Organizational Affiliation: