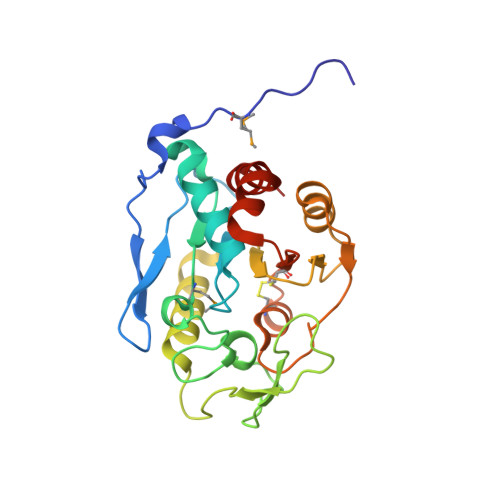
Mpaa is a Murein-Tripeptide-Specific Zinc Carboxypeptidase that Functions as Part of a Catabolic Pathway for Peptidoglycan Derived Peptides in Gamma-Proteobacteria.
Maqbool, A., Herve, M., Mengin-Lecreulx, D., Wilkinson, A.J., Thomas, G.H.(2012) Biochem J 448: 329
- PubMed: 22970852 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20121164
- Primary Citation Related Structures:
4AXV - PubMed Abstract:
The murein peptide amidase MpaA is a cytoplasmic enzyme that processes peptides derived from the turnover of murein. We have purified the enzyme from Escherichia coli and demonstrated that it efficiently hydrolyses the γ-D-glutamyl-diaminopimelic acid bond in the murein tripeptide (L-Ala-γ-D-Glu-meso-Dap), with Km and kcat values of 0.41±0.05 mM and 38.3±10 s-1. However, it is unable to act on the murein tetrapeptide (L-Ala-γ-D-Glu-meso-Dap-D-Ala). E. coli MpaA is a homodimer containing one bound zinc ion per chain, as judged by mass spectrometric analysis and size-exclusion chromatography. To investigate the structure of MpaA we solved the crystal structure of the orthologous protein from Vibrio harveyi to 2.17 Å (1Å=0.1 nm). Vh_MpaA, which has identical enzymatic and biophysical properties to the E. coli enzyme, has high structural similarity to eukaryotic zinc carboxypeptidases. The structure confirms that MpaA is a dimeric zinc metalloprotein. Comparison of the structure of MpaA with those of other carboxypeptidases reveals additional structure that partially occludes the substrate-binding groove, perhaps explaining the narrower substrate specificity of the enzyme compared with other zinc carboxypeptidases. In γ-proteobacteria mpaA is often located adjacent to mppA which encodes a periplasmic transporter protein previously shown to bind murein tripeptide. We demonstrate that MppA can also bind murein tetrapeptide with high affinity. The genetic coupling of these genes and their related biochemical functions suggest that MpaA amidase and MppA transporter form part of a catabolic pathway for utilization of murein-derived peptides that operates in γ-proteobacteria in addition to the established murein recycling pathways.
- Department of Biology (Area 10), Department of Chemistry, University of York, York YO10 5DD, UK.
Organizational Affiliation: