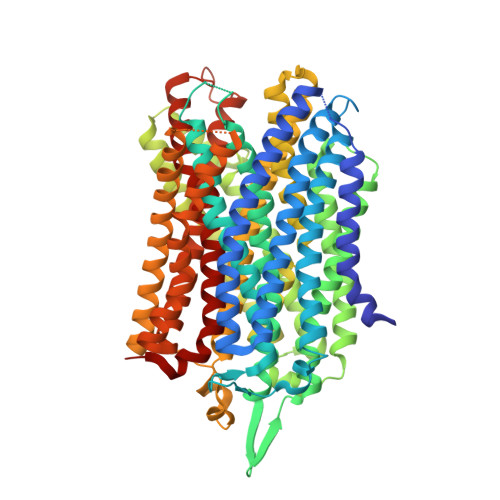
The Structure and Catalytic Cycle of a Sodium-Pumping Pyrophosphatase.
Kellosalo, J., Kajander, T., Kogan, K., Pokharel, K., Goldman, A.(2012) Science 337: 473
- PubMed: 22837527 Search on PubMed
- DOI: https://doi.org/10.1126/science.1222505
- Primary Citation Related Structures:
4AV3, 4AV6 - PubMed Abstract:
Membrane-integral pyrophosphatases (M-PPases) are crucial for the survival of plants, bacteria, and protozoan parasites. They couple pyrophosphate hydrolysis or synthesis to Na(+) or H(+) pumping. The 2.6-angstrom structure of Thermotoga maritima M-PPase in the resting state reveals a previously unknown solution for ion pumping. The hydrolytic center, 20 angstroms above the membrane, is coupled to the gate formed by the conserved Asp(243), Glu(246), and Lys(707) by an unusual "coupling funnel" of six α helices. Comparison with our 4.0-angstrom resolution structure of the product complex suggests that helix 12 slides down upon substrate binding to open the gate by a simple binding-change mechanism. Below the gate, four helices form the exit channel. Superimposing helices 3 to 6, 9 to 12, and 13 to 16 suggests that M-PPases arose through gene triplication.
- Structural Biology and Biophysics Program, Institute of Biotechnology, Post Office Box 65, University of Helsinki, FIN-00014, Finland.
Organizational Affiliation: