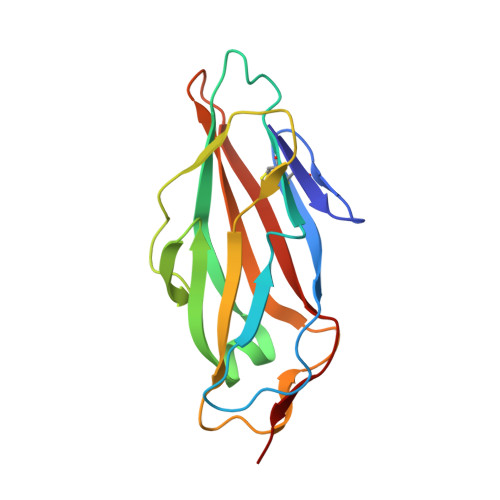
The Tyrosine Gate as a Potential Entropic Lever in the Receptor-Binding Site of the Bacterial Adhesin Fimh.
Wellens, A., Lahmann, M., Touaibia, M., Vaucher, J., Oscarson, S., Roy, R., Remaut, H., Bouckaert, J.(2012) Biochemistry 51: 4790
- PubMed: 22657089
- DOI: https://doi.org/10.1021/bi300251r
- Primary Citation Related Structures:
4AUU, 4AUY, 4AV0, 4AV4, 4AV5, 4AVH, 4AVI, 4AVJ, 4AVK - PubMed Abstract:
Uropathogenic Escherichia coli (UPEC) are the major causative agents of urinary tract infections. During infection, UPEC adhere to mannosylated glycoreceptors on the urothelium via the FimH adhesin located at the tip of type 1 pili. Synthetic FimH antiadhesives such as alkyl and phenyl α-D-mannopyranosides are thus ideal candidates for the chemical interception of this crucial step in pathogenesis. The crystal structures of the FimH lectin domain in its ligand-free form and in complexes with eight medium- and high-affinity mannopyranoside inhibitors are presented. The thermodynamic profiles of the FimH-inhibitor interactions indicate that the binding of FimH to α-D-mannopyranose is enthalpy-driven and has a negative entropic change. Addition of a hydrophobic aglycon influences the binding enthalpy and can induce a favorable entropic change. The alleviation of the entropic cost is at least in part explained by increased dynamics in the tyrosine gate (Tyr48 and Tyr137) of the FimH receptor-binding site upon binding of the ligand. Ligands with a phenyl group directly linked to the anomeric oxygen of α-D-mannose introduce the largest dynamics into the Tyr48 side chain, because conjugation with the anomeric oxygen of α-D-mannose forces the aromatic aglycon into a conformation that comes into close contact (≈2.65 Å) with Tyr48. A propargyl group in this position predetermines the orientation of the aglycon and significantly decreases affinity. FimH has the highest affinity for α-D-mannopyranosides substituted with hydrophobic aglycons that are compatible in shape and electrostatic properties to the tyrosine gate, such as heptyl α-D-mannose.
- Structural Molecular Microbiology, Vrije Universiteit Brussel, VIB, Brussels, Belgium.
Organizational Affiliation: