Synthesis and Structure-Activity Relationship of 4-(1,3-Benzothiazol-2-Yl)-Thiophene-2-Sulfonamides as Cyclin-Dependent Kinase 5 (Cdk5)/P25 Inhibitors.
Malmstrom, J., Viklund, J., Slivo, C., Costa, A., Maudet, M., Sandelin, C., Hiller, G., Olsson, L.L., Aagaard, A., Geschwindner, S., Xue, Y., Vasange, M.(2012) Bioorg Med Chem Lett 22: 5919
- PubMed: 22889803 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2012.07.068
- Primary Citation Related Structures:
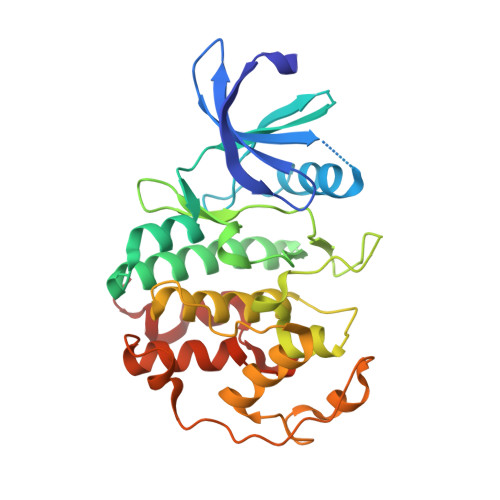
4AU8 - PubMed Abstract:
4-(1,3-Benzothiazol-2-yl)thiophene-2-sulfonamide (4a) was found to be a moderately potent inhibitor of cyclin-dependent kinase 5 (cdk5) from a HTS screen. The synthesis and SAR around this hit is described. The X-ray coordinates of ligand 4a with cdk5 are also reported, showing an unusual binding mode to the hinge region via a water molecule.
- Medicinal Chemistry, CNSP iMed Science, AstraZeneca R&D, Innovative Medicines, SE-15185 Södertälje, Sweden. Jonas.Malmstrom@astrazeneca.com
Organizational Affiliation: