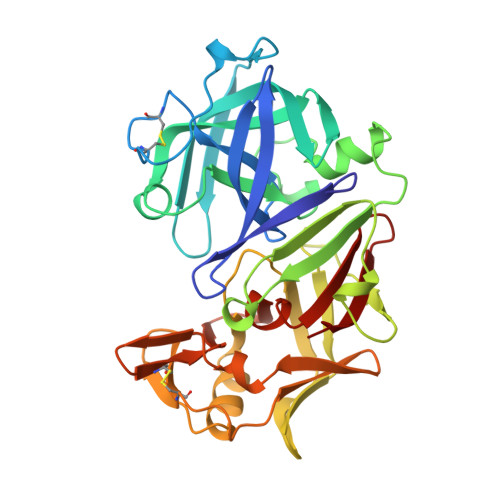
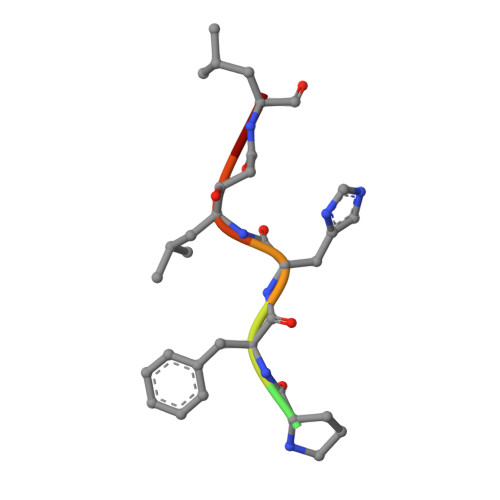
Structures of complexes of rhizopuspepsin with pepstatin and other statine-containing inhibitors.
Suguna, K., Padlan, E.A., Bott, R., Boger, J., Parris, K.D., Davies, D.R.(1992) Proteins 13: 195-205
- PubMed: 1603809 Search on PubMed
- DOI: https://doi.org/10.1002/prot.340130303
- Primary Citation Related Structures:
4APR, 5APR, 6APR - PubMed Abstract:
The three-dimensional structures of the complexes of the aspartic proteinase from Rhizopus chinensis (Rhizopuspepsin, EC 3.4.23.6) with pepstatin and two pepstatin-like peptide inhibitors of renin have been determined by X-ray diffraction methods and refined by restrained least-squares procedures. The inhibitors adopt an extended conformation and lie in the deep groove located between the two domains of the enzyme. Inhibitor binding is accompanied by a conformational change at the "flap," a beta-hairpin loop region, that projects over the binding cleft and closes down over the inhibitor, excluding water molecules from the vicinity of the scissile bond. The hydroxyl group of the central statyl residue of the inhibitors replaces the water molecule found between the two active aspartates, Asp-35 and Asp-218, in the native structure. The refined structures provide additional data to define the specific subsites of the enzyme and also show a system of hydrogen bonding to the inhibitor backbone similar to that observed for a reduced inhibitor.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892.
Organizational Affiliation: